Building a Quant Trading Pipeline: Deep Learning vs. Classical Machine Learning
Evaluating Volume, Volatility, and Return Features with Walk-Forward Cross-Validation
Download Source code using the button at the end of this article!
Financial markets are notoriously noisy, making directional forecasting—predicting whether the S&P 500 will be up or down a month from now—one of the most challenging problems in algorithmic trading. Many machine learning models look incredible in a backtest but fail miserably in live markets because they accidentally peek into the future during training. To build a true quantitative trading edge, you need more than just a complex algorithm; you need a bulletproof validation strategy that strictly respects the chronological flow of time.
In this walkthrough, we are going to build a complete, end-to-end quantitative machine learning pipeline from the ground up. We will engineer multi-horizon market signals—scaling from simple daily log returns to 80-week rolling volatility and trading volume—and test them using a rigorous walk-forward cross-validation approach to eliminate data leakage. Then, we will set up a head-to-head benchmarking experiment: pitting a highly regularized classical model (Elastic Net Logistic Regression) directly against a recurrent deep learning architecture (a stateful LSTM) to see which approach best captures underlying market dynamics.
By the end of this article, you will understand how to systematically isolate and evaluate different feature sets to determine which data points actually carry predictive power. You can expect a practical deep dive into hyperparameter tuning, structuring temporal data arrays for Keras and Scikit-Learn, and rigorously grading your backtest results using precision, recall, and ROC-AUC curves. Whether you are a data scientist stepping into finance or a quant looking to refine your modeling workflow, this guide will give you a reusable framework for testing your own algorithmic trading signals.
#Standard libraries
import pandas as pd
import seaborn as sns
import numpy as np
import matplotlib.pyplot as plt
import time
# library for sampling
from scipy.stats import uniform
# libraries for Data Download
import datetime
from pandas_datareader import data as pdr
import fix_yahoo_finance as yf
# sklearn
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import TimeSeriesSplit
from sklearn.model_selection import RandomizedSearchCV
from sklearn.model_selection import GridSearchCV
from sklearn import metrics
from sklearn.metrics import make_scorer
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn import linear_model
# Keras
import keras
from keras.models import Sequential
from keras.layers import LSTM
from keras.layers import Dense
from keras.layers import Dropout
from keras.wrappers.scikit_learn import KerasClassifierOutput
[stderr]
Using TensorFlow backend.Importing pandas as pd brings the DataFrame/Seriesâcentric API that the notebook relies on to ingest, align and manipulate the historical price series and engineered features; youâll see it used for reading and writing tabular data, handling timestamps and indices, performing joins/concatenations, shifting/rolling/resampling for lagged and windowed features, filling or interpolating missing values, and slicing the timeâconsistent train/test partitions that are later converted into LSTM input arrays. Because the experimentâs pipeline frames raw price histories into supervised, windowed feature matrices and normalizes and partitions them, pandas is the foundational tool used throughout the preprocessing and feature engineering stages. The repeated start=time.time() lines found elsewhere serve a different purpose: they are adâhoc runtime timers wrapped around model training blocks to report elapsed time, whereas pandas is a core data manipulation dependency used across the entire data preparation workflow.
2. Create Classes
# Define a callback class
# Resets the states after each epoch (after going through a full time series)
class ModelStateReset(keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs={}):
self.model.reset_states()
reset=ModelStateReset()
# Different Approach
#class modLSTM(LSTM):
# def call(self, x, mask=None):
# if self.stateful:
# self.reset_states()
# return super(modLSTM, self).call(x, mask)ModelStateReset is a small Keras callback class used by the notebook to explicitly clear the LSTM’s recurrent state at the end of every training epoch. In this pipeline the LSTM created by create_shallow_LSTM is configured to keep state across batches to model temporal dependencies, and training is performed with time-ordered cross-validation and shuffle disabled, so internal hidden and cell states would otherwise persist across epoch boundaries or between CV folds and could leak information. Keras calls the callback’s on_epoch_end hook automatically at the end of each epoch, and that hook triggers the LSTM’s reset_states method so the next epoch or fold starts from a clean recurrent state. The notebook wires an instance named reset into the fit call (so GridSearchCV/KerasClassifier training will execute it), and the commented alternative shows a different approach that would subclass the LSTM layer itself to reset states during its call rather than relying on a training callback; the callback approach keeps the state-management logic separate from the model construction and is the mechanism actually used in the comparisons between recurrent and classical models.
3. Write Functions
In [3] Copy
# Function to create an LSTM model, required for KerasClassifier
def create_shallow_LSTM(epochs=1,
LSTM_units=1,
num_samples=1,
look_back=1,
num_features=None,
dropout_rate=0,
recurrent_dropout=0,
verbose=0):
model=Sequential()
model.add(LSTM(units=LSTM_units,
batch_input_shape=(num_samples, look_back, num_features),
stateful=True,
recurrent_dropout=recurrent_dropout))
model.add(Dropout(dropout_rate))
model.add(Dense(1, activation='sigmoid', kernel_initializer=keras.initializers.he_normal(seed=1)))
model.compile(loss='binary_crossentropy', optimizer="adam", metrics=['accuracy'])
return modelcreate_shallow_LSTM constructs and returns a ready-to-train Keras Sequential model that the experimental pipeline wraps with KerasClassifier for hyperparameter search and cross-validated training. The function builds a shallow architecture: a single LSTM layer configured to be stateful and to receive a fixed batch input shape determined by the supplied num_samples, look_back and num_features so that training examples are treated as short time-windowed sequences consistent with how the preprocessing frames the series; the LSTM layer size is controlled by LSTM_units and it applies recurrent dropout to regularize the recurrent connections. A Dropout layer follows to regularize the LSTM outputs, and a final Dense output layer with a single sigmoid unit produces a probability for the binary up/down target used in the forecasting experiments; the Dense kernel is initialized with a He normal initializer seeded for reproducibility. The model is compiled with binary cross-entropy loss, the Adam optimizer and accuracy as a tracked metric so it can be evaluated by GridSearchCV using the provided scoring. Because the model is stateful and requires a fixed batch configuration, create_shallow_LSTM accepts the batch-related parameters that the surrounding code supplies (and the GridSearchCV wrappers vary batch_size during tuning), and then returns the compiled model for the rest of the pipeline to fit and evaluate.
4. Data
4.1 Import Raw Data
In [4] Copy
# Imports data
start_sp=datetime.datetime(1980, 1, 1)
end_sp=datetime.datetime(2019, 2, 28)
yf.pdr_override()
sp500=pdr.get_data_yahoo('^GSPC',
start_sp,
end_sp)
sp500.shapeOutput
[stdout]
[*********************100%***********************] 1 of 1 downloaded(9875, 6)start_sp is a datetime object that sets the lower bound of the historical S&P 500 series the notebook pulls from Yahoo Finance; it and the companion end_sp define the exact temporal window that pandas_datareader (after calling yf.pdr_override) will fetch into the sp500 DataFrame. Choosing 1980-01-01 deliberately supplies a long, multiâdecade record so the downstream feature engineeringâsuch as creating Log_Ret_1d and many long rolling-sum return featuresâhas enough prehistory to compute large-window statistics (the notebook computes rolling windows up to several hundred trading days) and so the LSTM framing can produce many time-windowed sequences while preserving a temporally consistent train/test split. Because the same sp500 object is subsequently used for plotting and model training, the start_sp value directly controls the sample available for normalization, cross-validation across time, and the experimental comparisons between recurrent and classical learners, and it therefore anchors reproducibility of the endâtoâend pipeline.
4.2 Create Features
In [5] Copy
# Compute the logarithmic returns using the Closing price
sp500['Log_Ret_1d']=np.log(sp500['Close'] / sp500['Close'].shift(1))
# Compute logarithmic returns using the pandas rolling mean function
sp500['Log_Ret_1w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=5).sum()
sp500['Log_Ret_2w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=10).sum()
sp500['Log_Ret_3w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=15).sum()
sp500['Log_Ret_4w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=20).sum()
sp500['Log_Ret_8w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=40).sum()
sp500['Log_Ret_12w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=60).sum()
sp500['Log_Ret_16w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=80).sum()
sp500['Log_Ret_20w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=100).sum()
sp500['Log_Ret_24w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=120).sum()
sp500['Log_Ret_28w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=140).sum()
sp500['Log_Ret_32w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=160).sum()
sp500['Log_Ret_36w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=180).sum()
sp500['Log_Ret_40w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=200).sum()
sp500['Log_Ret_44w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=220).sum()
sp500['Log_Ret_48w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=240).sum()
sp500['Log_Ret_52w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=260).sum()
sp500['Log_Ret_56w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=280).sum()
sp500['Log_Ret_60w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=300).sum()
sp500['Log_Ret_64w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=320).sum()
sp500['Log_Ret_68w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=340).sum()
sp500['Log_Ret_72w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=360).sum()
sp500['Log_Ret_76w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=380).sum()
sp500['Log_Ret_80w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=400).sum()
# Compute Volatility using the pandas rolling standard deviation function
sp500['Vol_1w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=5).std()*np.sqrt(5)
sp500['Vol_2w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=10).std()*np.sqrt(10)
sp500['Vol_3w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=15).std()*np.sqrt(15)
sp500['Vol_4w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=20).std()*np.sqrt(20)
sp500['Vol_8w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=40).std()*np.sqrt(40)
sp500['Vol_12w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=60).std()*np.sqrt(60)
sp500['Vol_16w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=80).std()*np.sqrt(80)
sp500['Vol_20w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=100).std()*np.sqrt(100)
sp500['Vol_24w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=120).std()*np.sqrt(120)
sp500['Vol_28w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=140).std()*np.sqrt(140)
sp500['Vol_32w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=160).std()*np.sqrt(160)
sp500['Vol_36w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=180).std()*np.sqrt(180)
sp500['Vol_40w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=200).std()*np.sqrt(200)
sp500['Vol_44w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=220).std()*np.sqrt(220)
sp500['Vol_48w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=240).std()*np.sqrt(240)
sp500['Vol_52w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=260).std()*np.sqrt(260)
sp500['Vol_56w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=280).std()*np.sqrt(280)
sp500['Vol_60w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=300).std()*np.sqrt(300)
sp500['Vol_64w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=320).std()*np.sqrt(320)
sp500['Vol_68w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=340).std()*np.sqrt(340)
sp500['Vol_72w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=360).std()*np.sqrt(360)
sp500['Vol_76w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=380).std()*np.sqrt(380)
sp500['Vol_80w']=pd.Series(sp500['Log_Ret_1d']).rolling(window=400).std()*np.sqrt(400)
# Compute Volumes using the pandas rolling mean function
sp500['Volume_1w']=pd.Series(sp500['Volume']).rolling(window=5).mean()
sp500['Volume_2w']=pd.Series(sp500['Volume']).rolling(window=10).mean()
sp500['Volume_3w']=pd.Series(sp500['Volume']).rolling(window=15).mean()
sp500['Volume_4w']=pd.Series(sp500['Volume']).rolling(window=20).mean()
sp500['Volume_8w']=pd.Series(sp500['Volume']).rolling(window=40).mean()
sp500['Volume_12w']=pd.Series(sp500['Volume']).rolling(window=60).mean()
sp500['Volume_16w']=pd.Series(sp500['Volume']).rolling(window=80).mean()
sp500['Volume_20w']=pd.Series(sp500['Volume']).rolling(window=100).mean()
sp500['Volume_24w']=pd.Series(sp500['Volume']).rolling(window=120).mean()
sp500['Volume_28w']=pd.Series(sp500['Volume']).rolling(window=140).mean()
sp500['Volume_32w']=pd.Series(sp500['Volume']).rolling(window=160).mean()
sp500['Volume_36w']=pd.Series(sp500['Volume']).rolling(window=180).mean()
sp500['Volume_40w']=pd.Series(sp500['Volume']).rolling(window=200).mean()
sp500['Volume_44w']=pd.Series(sp500['Volume']).rolling(window=220).mean()
sp500['Volume_48w']=pd.Series(sp500['Volume']).rolling(window=240).mean()
sp500['Volume_52w']=pd.Series(sp500['Volume']).rolling(window=260).mean()
sp500['Volume_56w']=pd.Series(sp500['Volume']).rolling(window=280).mean()
sp500['Volume_60w']=pd.Series(sp500['Volume']).rolling(window=300).mean()
sp500['Volume_64w']=pd.Series(sp500['Volume']).rolling(window=320).mean()
sp500['Volume_68w']=pd.Series(sp500['Volume']).rolling(window=340).mean()
sp500['Volume_72w']=pd.Series(sp500['Volume']).rolling(window=360).mean()
sp500['Volume_76w']=pd.Series(sp500['Volume']).rolling(window=380).mean()
sp500['Volume_80w']=pd.Series(sp500['Volume']).rolling(window=400).mean()
# Label data: Up (Down) if the the 1 month (≈ 21 trading days) logarithmic return increased (decreased)
sp500['Return_Label']=pd.Series(sp500['Log_Ret_1d']).shift(-21).rolling(window=21).sum()
sp500['Label']=np.where(sp500['Return_Label'] > 0, 1, 0)
# Drop NA´s
sp500=sp500.dropna("index")
sp500=sp500.drop(['Open', 'High', 'Low', 'Close', 'Adj Close', 'Volume', "Return_Label"], axis=1)The statement that creates Log_Ret_1d computes the oneâday logarithmic return from the closing price series in sp500 by taking the natural logarithm of the ratio between todayâs Close and the previous dayâs Close (using pandas shift to reference the prior row), producing an additive, stationary return series that the rest of the feature engineering operates on. That oneâday log return is then aggregated with pandas rolling and sum to produce multiâhorizon log returns (named Log_Ret_1w through Log_Ret_80w) so each longerâhorizon return is just the sum of daily log returns over the window, and the same daily series is used with rolling and std (scaled by the square root of the window length) to produce timeâscale volatility features (Vol_1w through Vol_80w). The Volume_* features are made by applying pandas rolling mean to the raw Volume column across the same windows. For supervised labeling, the code shifts the oneâday log returns forward by roughly 21 trading days and sums over a 21âday rolling window to compute the approximate oneâmonth forward return (Return_Label), then binarizes that into Label with numpy where to indicate whether the forward return is positive. Missing values produced by shift and rolling are removed with pandas dropna, and the original raw price and intermediate columns are dropped so the resulting sp500 frame contains only the engineered return, volatility, volume and label columns. These engineered columns are the inputs referenced later by train_test_split and the iloc slices used for plotting and for assembling baseline and LSTM model feature sets, so Log_Ret_1d is the foundational time series from which the multiâscale features and the supervised target are derived.
4.3 Extract Basic Information
In [6] Copy
# Show rows and columns
print("Rows, Columns:");print(sp500.shape);print("\n")
# Describe DataFrame columns
print("Columns:");print(sp500.columns);print("\n")
# Show info on DataFrame
print("Info:");print(sp500.info()); print("\n")
# Count Non-NA values
print("Non-NA:");print(sp500.count()); print("\n")
# Show head
print("Head");print(sp500.head()); print("\n")
# Show tail
print("Tail");print(sp500.tail());print("\n")
# Show summary statistics
print("Summary statistics:");print(sp500.describe());print("\n")Output
[stdout]
Rows, Columns:
(9454, 71)
Columns:
Index([’Log_Ret_1d’, ‘Log_Ret_1w’, ‘Log_Ret_2w’, ‘Log_Ret_3w’, ‘Log_Ret_4w’,
‘Log_Ret_8w’, ‘Log_Ret_12w’, ‘Log_Ret_16w’, ‘Log_Ret_20w’,
‘Log_Ret_24w’, ‘Log_Ret_28w’, ‘Log_Ret_32w’, ‘Log_Ret_36w’,
‘Log_Ret_40w’, ‘Log_Ret_44w’, ‘Log_Ret_48w’, ‘Log_Ret_52w’,
‘Log_Ret_56w’, ‘Log_Ret_60w’, ‘Log_Ret_64w’, ‘Log_Ret_68w’,
‘Log_Ret_72w’, ‘Log_Ret_76w’, ‘Log_Ret_80w’, ‘Vol_1w’, ‘Vol_2w’,
‘Vol_3w’, ‘Vol_4w’, ‘Vol_8w’, ‘Vol_12w’, ‘Vol_16w’, ‘Vol_20w’,
‘Vol_24w’, ‘Vol_28w’, ‘Vol_32w’, ‘Vol_36w’, ‘Vol_40w’, ‘Vol_44w’,
‘Vol_48w’, ‘Vol_52w’, ‘Vol_56w’, ‘Vol_60w’, ‘Vol_64w’, ‘Vol_68w’,
‘Vol_72w’, ‘Vol_76w’, ‘Vol_80w’, ‘Volume_1w’, ‘Volume_2w’, ‘Volume_3w’,
‘Volume_4w’, ‘Volume_8w’, ‘Volume_12w’, ‘Volume_16w’, ‘Volume_20w’,
‘Volume_24w’, ‘Volume_28w’, ‘Volume_32w’, ‘Volume_36w’, ‘Volume_40w’,
‘Volume_44w’, ‘Volume_48w’, ‘Volume_52w’, ‘Volume_56w’, ‘Volume_60w’,
‘Volume_64w’, ‘Volume_68w’, ‘Volume_72w’, ‘Volume_76w’, ‘Volume_80w’,
‘Label’],
dtype=’object’)
Info:
<class ‘pandas.core.frame.DataFrame’>
DatetimeIndex: 9454 entries, 1981-07-31 to 2019-01-28
Data columns (total 71 columns):
Log_Ret_1d 9454 non-null float64
Log_Ret_1w 9454 non-null float64
Log_Ret_2w 9454 non-null float64
Log_Ret_3w 9454 non-null float64
Log_Ret_4w 9454 non-null float64
Log_Ret_8w 9454 non-null float64
Log_Ret_12w 9454 non-null float64
Log_Ret_16w 9454 non-null float64
Log_Ret_20w 9454 non-null float64
Log_Ret_24w 9454 non-null float64
Log_Ret_28w 9454 non-null float64
Log_Ret_32w 9454 non-null float64
Log_Ret_36w 9454 non-null float64
Log_Ret_40w 9454 non-null float64
Log_Ret_44w 9454 non-null float64
Log_Ret_48w 9454 non-null float64
Log_Ret_52w 9454 non-null float64
Log_Ret_56w 9454 non-null float64
Log_Ret_60w 9454 non-null float64
Log_Ret_64w 9454 non-null float64
Log_Ret_68w 9454 non-null float64
Log_Ret_72w 9454 non-null float64
Log_Ret_76w 9454 non-null float64
Log_Ret_80w 9454 non-null float64
Vol_1w 9454 non-null float64
Vol_2w 9454 non-null float64
Vol_3w 9454 non-null float64
Vol_4w 9454 non-null float64
Vol_8w 9454 non-null float64
Vol_12w 9454 non-null float64
Vol_16w 9454 non-null float64
Vol_20w 9454 non-null float64
Vol_24w 9454 non-null float64
Vol_28w 9454 non-null float64
Vol_32w 9454 non-null float64
Vol_36w 9454 non-null float64
Vol_40w 9454 non-null float64
Vol_44w 9454 non-null float64
Vol_48w 9454 non-null float64
Vol_52w 9454 non-null float64
Vol_56w 9454 non-null float64
Vol_60w 9454 non-null float64
Vol_64w 9454 non-null float64
Vol_68w 9454 non-null float64
Vol_72w 9454 non-null float64
Vol_76w 9454 non-null float64
Vol_80w 9454 non-null float64
Volume_1w 9454 non-null float64
Volume_2w 9454 non-null float64
Volume_3w 9454 non-null float64
Volume_4w 9454 non-null float64
Volume_8w 9454 non-null float64
Volume_12w 9454 non-null float64
Volume_16w 9454 non-null float64
Volume_20w 9454 non-null float64
Volume_24w 9454 non-null float64
Volume_28w 9454 non-null float64
Volume_32w 9454 non-null float64
Volume_36w 9454 non-null float64
Volume_40w 9454 non-null float64
Volume_44w 9454 non-null float64
Volume_48w 9454 non-null float64
Volume_52w 9454 non-null float64
Volume_56w 9454 non-null float64
Volume_60w 9454 non-null float64
Volume_64w 9454 non-null float64
Volume_68w 9454 non-null float64
Volume_72w 9454 non-null float64
Volume_76w 9454 non-null float64
Volume_80w 9454 non-null float64
Label 9454 non-null int64
dtypes: float64(70), int64(1)
memory usage: 5.2 MB
None
Non-NA:
Log_Ret_1d 9454
Log_Ret_1w 9454
Log_Ret_2w 9454
Log_Ret_3w 9454
Log_Ret_4w 9454
Log_Ret_8w 9454
Log_Ret_12w 9454
Log_Ret_16w 9454
Log_Ret_20w 9454
Log_Ret_24w 9454
Log_Ret_28w 9454
Log_Ret_32w 9454
Log_Ret_36w 9454
Log_Ret_40w 9454
Log_Ret_44w 9454
Log_Ret_48w 9454
Log_Ret_52w 9454
Log_Ret_56w 9454
Log_Ret_60w 9454
Log_Ret_64w 9454
Log_Ret_68w 9454
Log_Ret_72w 9454
Log_Ret_76w 9454
Log_Ret_80w 9454
Vol_1w 9454
Vol_2w 9454
Vol_3w 9454
Vol_4w 9454
Vol_8w 9454
Vol_12w 9454
...
Vol_60w 9454
Vol_64w 9454
Vol_68w 9454
Vol_72w 9454
Vol_76w 9454
Vol_80w 9454
Volume_1w 9454
Volume_2w 9454
Volume_3w 9454
Volume_4w 9454
Volume_8w 9454
Volume_12w 9454
Volume_16w 9454
Volume_20w 9454
Volume_24w 9454
Volume_28w 9454
Volume_32w 9454
Volume_36w 9454
Volume_40w 9454
Volume_44w 9454
Volume_48w 9454
Volume_52w 9454
Volume_56w 9454
Volume_60w 9454
Volume_64w 9454
Volume_68w 9454
Volume_72w 9454
Volume_76w 9454
Volume_80w 9454
Label 9454
Length: 71, dtype: int64
Head
Log_Ret_1d Log_Ret_1w Log_Ret_2w Log_Ret_3w Log_Ret_4w \
Date
1981-07-31 0.006975 0.018969 0.001223 0.011910 0.017569
1981-08-03 -0.003367 0.004455 0.013580 0.006459 0.024124
1981-08-04 0.005350 0.015673 0.021887 0.011732 0.022667
1981-08-05 0.011294 0.026813 0.042655 0.018563 0.033338
1981-08-06 -0.000226 0.020027 0.040307 0.017492 0.025503
Log_Ret_8w Log_Ret_12w Log_Ret_16w Log_Ret_20w Log_Ret_24w \
Date
1981-07-31 -0.000306 0.001070 -0.022582 0.003520 0.012683
1981-08-03 -0.013247 -0.009079 -0.028931 0.004070 0.009549
1981-08-04 -0.008048 -0.003652 -0.026257 -0.015206 0.022667
1981-08-05 0.005290 0.022564 -0.013774 -0.003311 0.039905
1981-08-06 0.002415 0.014581 -0.003838 -0.015263 0.043609
... Volume_48w Volume_52w Volume_56w Volume_60w \
Date ...
1981-07-31 ... 4.800596e+07 4.795673e+07 4.766939e+07 4.717517e+07
1981-08-03 ... 4.799646e+07 4.790835e+07 4.768893e+07 4.715470e+07
1981-08-04 ... 4.798354e+07 4.788362e+07 4.769511e+07 4.715020e+07
1981-08-05 ... 4.799821e+07 4.792927e+07 4.772293e+07 4.720257e+07
1981-08-06 ... 4.797262e+07 4.799012e+07 4.774779e+07 4.723613e+07
Volume_64w Volume_68w Volume_72w Volume_76w Volume_80w \
Date
1981-07-31 46354531.25 4.569368e+07 4.540333e+07 4.563068e+07 45952075.0
1981-08-03 46389093.75 4.562300e+07 4.540144e+07 4.560037e+07 45949675.0
1981-08-04 46416781.25 4.560165e+07 4.540325e+07 4.553079e+07 45922125.0
1981-08-05 46499125.00 4.565591e+07 4.544658e+07 4.555100e+07 45960025.0
1981-08-06 46565437.50 4.571426e+07 4.546814e+07 4.557468e+07 45978950.0
Label
Date
1981-07-31 0
1981-08-03 0
1981-08-04 0
1981-08-05 0
1981-08-06 0
[5 rows x 71 columns]
Tail
Log_Ret_1d Log_Ret_1w Log_Ret_2w Log_Ret_3w Log_Ret_4w \
Date
2019-01-22 -0.014258 0.019285 0.032114 0.057515 0.064913
2019-01-23 0.002200 0.010821 0.024666 0.051259 0.087916
2019-01-24 0.001375 0.009976 0.021951 0.051366 0.116778
2019-01-25 0.008453 0.010867 0.025896 0.084888 0.076827
2019-01-28 -0.007878 -0.010108 0.018164 0.043250 0.060423
Log_Ret_8w Log_Ret_12w Log_Ret_16w Log_Ret_20w Log_Ret_24w \
Date
2019-01-22 -0.003409 -0.040124 -0.101976 -0.095500 -0.062462
2019-01-23 -0.004247 -0.006573 -0.096481 -0.093569 -0.065134
2019-01-24 0.003704 -0.023652 -0.097866 -0.097879 -0.062718
2019-01-25 -0.003256 0.002281 -0.089406 -0.084986 -0.059180
2019-01-28 -0.014390 0.000984 -0.100918 -0.092999 -0.071691
... Volume_48w Volume_52w Volume_56w Volume_60w \
Date ...
2019-01-22 ... 3.601485e+09 3.629041e+09 3.599487e+09 3.587928e+09
2019-01-23 ... 3.596106e+09 3.628588e+09 3.600307e+09 3.586275e+09
2019-01-24 ... 3.588305e+09 3.628038e+09 3.601526e+09 3.586096e+09
2019-01-25 ... 3.580530e+09 3.628702e+09 3.602449e+09 3.587466e+09
2019-01-28 ... 3.578684e+09 3.628852e+09 3.602700e+09 3.587370e+09
Volume_64w Volume_68w Volume_72w Volume_76w \
Date
2019-01-22 3.582160e+09 3.559243e+09 3.530446e+09 3.520048e+09
2019-01-23 3.582736e+09 3.559011e+09 3.531507e+09 3.520774e+09
2019-01-24 3.583622e+09 3.554835e+09 3.532314e+09 3.521433e+09
2019-01-25 3.586429e+09 3.556658e+09 3.533421e+09 3.523418e+09
2019-01-28 3.588690e+09 3.557728e+09 3.535712e+09 3.525004e+09
Volume_80w Label
Date
2019-01-22 3.508435e+09 1
2019-01-23 3.508233e+09 1
2019-01-24 3.507829e+09 1
2019-01-25 3.508694e+09 1
2019-01-28 3.504530e+09 1
[5 rows x 71 columns]
Summary statistics:[stdout]
Log_Ret_1d Log_Ret_1w Log_Ret_2w Log_Ret_3w Log_Ret_4w \
count 9454.000000 9454.000000 9454.000000 9454.000000 9454.000000
mean 0.000319 0.001596 0.003191 0.004772 0.006342
std 0.011115 0.023453 0.031869 0.038598 0.044166
min -0.228997 -0.319214 -0.377868 -0.365360 -0.350374
25% -0.004497 -0.010109 -0.012040 -0.013111 -0.015094
50% 0.000531 0.003047 0.005691 0.008145 0.010847
75% 0.005554 0.014624 0.020748 0.026528 0.031437
max 0.109572 0.174887 0.195882 0.199030 0.211030
Log_Ret_8w Log_Ret_12w Log_Ret_16w Log_Ret_20w Log_Ret_24w \
count 9454.000000 9454.000000 9454.000000 9454.000000 9454.000000
mean 0.012641 0.019043 0.025468 0.031968 0.038525
std 0.061672 0.075067 0.086494 0.097284 0.108054
min -0.474377 -0.532590 -0.534617 -0.535123 -0.605207
25% -0.014885 -0.013295 -0.013128 -0.010956 -0.009207
50% 0.018085 0.026825 0.033895 0.040462 0.050485
75% 0.049131 0.063615 0.077685 0.090158 0.101540
max 0.289631 0.328761 0.325124 0.377440 0.421288
... Volume_48w Volume_52w Volume_56w Volume_60w \
count ... 9.454000e+03 9.454000e+03 9.454000e+03 9.454000e+03
mean ... 1.642717e+09 1.639007e+09 1.635288e+09 1.631596e+09
std ... 1.691191e+09 1.689579e+09 1.687964e+09 1.686398e+09
min ... 4.668229e+07 4.653250e+07 4.665168e+07 4.691313e+07
25% ... 1.756674e+08 1.761194e+08 1.760642e+08 1.754544e+08
50% ... 8.858820e+08 8.861620e+08 8.787942e+08 8.702093e+08
75% ... 3.402212e+09 3.392543e+09 3.391066e+09 3.388343e+09
max ... 6.128274e+09 6.073674e+09 5.995085e+09 5.949259e+09
Volume_64w Volume_68w Volume_72w Volume_76w Volume_80w \
count 9.454000e+03 9.454000e+03 9.454000e+03 9.454000e+03 9.454000e+03
mean 1.627909e+09 1.624251e+09 1.620625e+09 1.617043e+09 1.613479e+09
std 1.684841e+09 1.683312e+09 1.681791e+09 1.680278e+09 1.678765e+09
min 4.635453e+07 4.560165e+07 4.540144e+07 4.529534e+07 4.521465e+07
25% 1.745719e+08 1.736218e+08 1.729558e+08 1.730130e+08 1.727768e+08
50% 8.689982e+08 8.589569e+08 8.484074e+08 8.447265e+08 8.424810e+08
75% 3.389822e+09 3.391340e+09 3.392891e+09 3.392742e+09 3.388016e+09
max 5.871179e+09 5.828395e+09 5.766422e+09 5.691528e+09 5.615789e+09
Label
count 9454.000000
mean 0.626296
std 0.483812
min 0.000000
25% 0.000000
50% 1.000000
75% 1.000000
max 1.000000
[8 rows x 71 columns]This cell performs a quick, textual sanity check of the sp500 DataFrame that was fetched earlier by pandas_datareader (the Yahoo reader described previously). It first inspects the DataFrameâs shape so you know the number of time steps and the number of columns coming out of the import and initial feature engineering (for example the Log_Ret_1d column you already saw created will be one of these columns). It then lists the DataFrameâs column names so you can verify which raw OHLCV fields and engineered features are present, and calls the DataFrame.info method to report dtypes and nonânull counts to reveal any missing values or unexpected types that would affect downstream normalization and windowing. The count method is used as a columnâwise nonâNA tally to corroborate the info output, head and tail show the first and last rows to confirm chronological ordering and boundary values consistent with the start_sp/end_sp range, and describe produces summary statistics (mean, std, min/max, percentiles) to give a quick view of feature distributions before framing the series into time windows and training models. This is similar to the earlier use of the shape attribute during data import but extends that single check into a fuller set of exploratory checks (textual rather than plotting) to catch problems early in the pipeline.
4.4 Plot Data
In [7] Copy
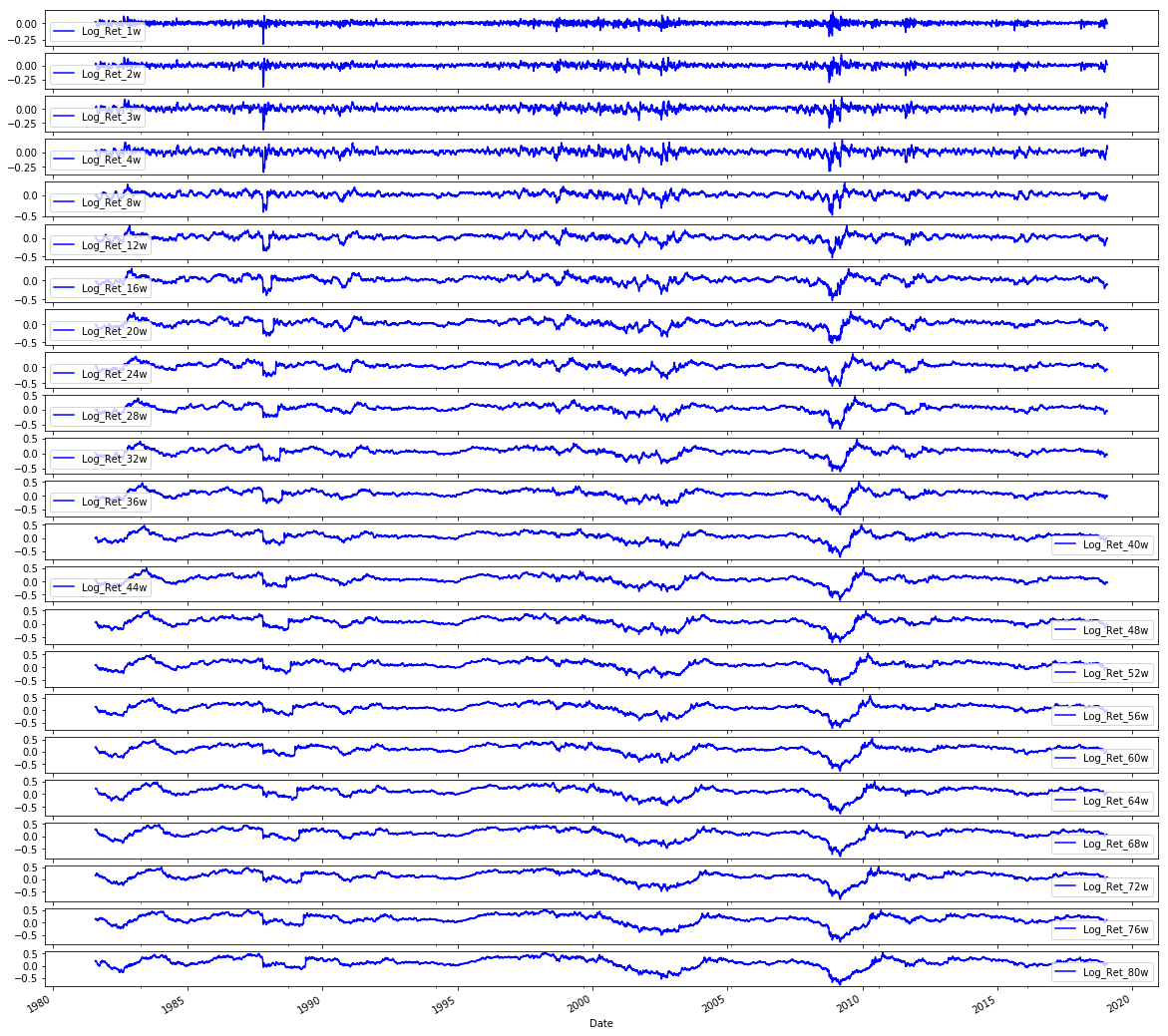
# Plot the logarithmic returns
sp500.iloc[:,1:24].plot(subplots=True, color='blue', figsize=(20, 20));Output
In the exploratory plotting step labeled 4.4 Plot Data the notebook uses pandas integer-location indexing via sp500.iloc to slice the set of engineered return-related columns (the block of columns immediately after the first column) and then invokes the DataFrame plotting facility to render each selected column as its own subplot. Because the selection is done with integer-location indexing, the time index from the sp500 DataFrame is preserved on the x-axis and each column becomes a separate vertical panel so you can compare series by eye; the call requests a large square figure and a uniform blue line color to make the many small series readable. In the pipeline this visualization is a sanity check on the feature engineering stage (remember the Log_Ret_1d column we created earlier is included in this slice) so you can inspect stationarity, volatility clustering, outliers and any preprocessing artifacts before framing windows and training models. The two similar snippets for volatilities and volumes follow the same pattern but operate on different contiguous column ranges (the volatility block and the volume block respectively), and the correlation-matrix code builds on a wider slice (returns plus volatilities) to compute and mask pairwise correlations for a focused heatmap comparison rather than plotting raw time series.
In [8] Copy
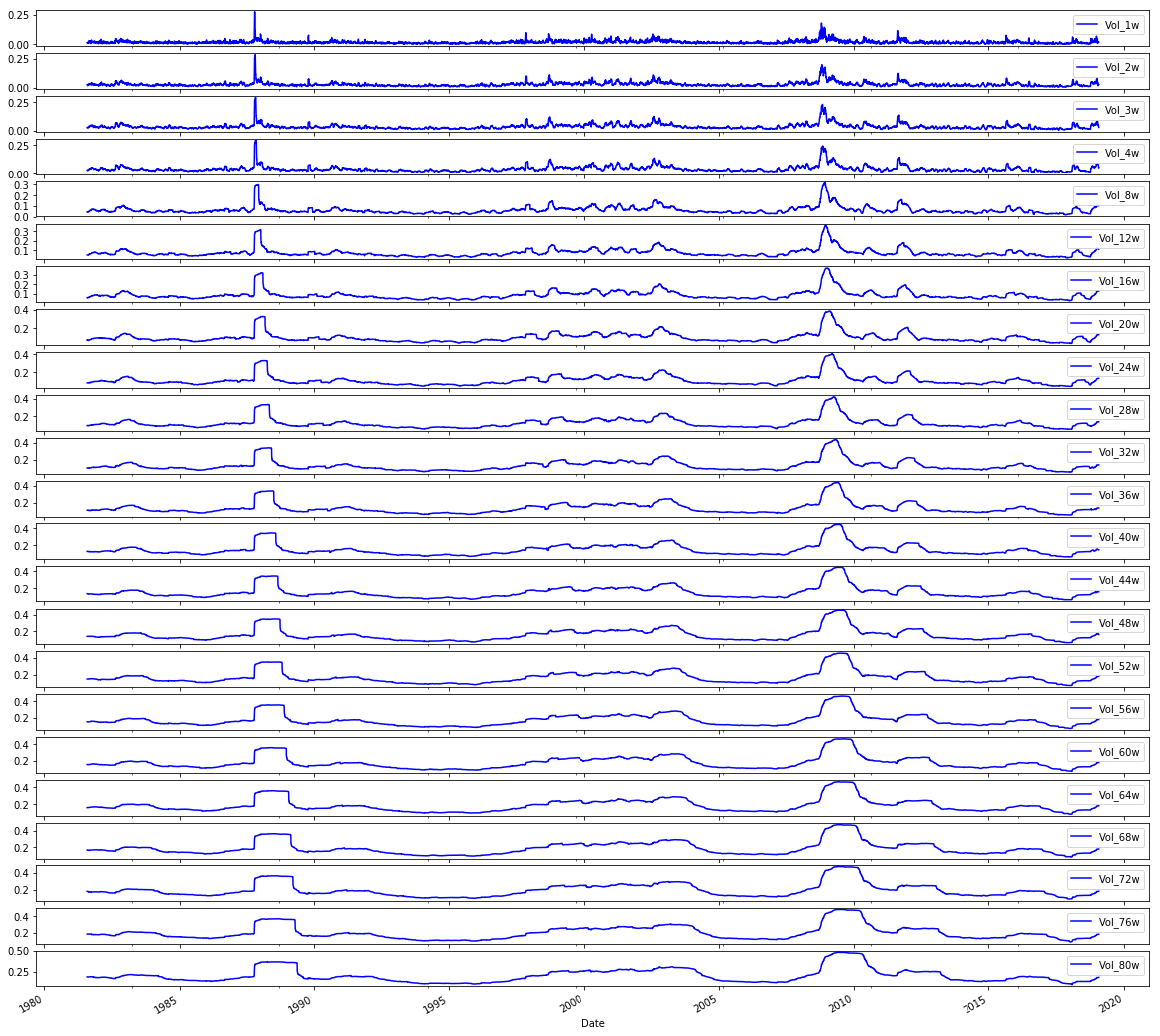
# Plot the Volatilities
sp500.iloc[:,24:47].plot(subplots=True, color='blue',figsize=(20, 20));Output
Here the notebook takes the preassembled sp500 DataFrame (the raw OHLCV series that has already been augmented with engineered features like the log returns and volatility metrics) and uses pandasâ integer-location indexing to grab the block of columns that represent the volatility features, then invokes the DataFrame plotting facility to render each volatility series into its own subplot with a uniform visual style (blue lines) on a large canvas so you can inspect each series independently. In the pipeline this is an exploratory visualization step that sits between feature engineering and the framing/normalization steps used to build LSTM sequences: its purpose is to let you visually validate the volatility feature set for trends, outliers, seasonal structure or data problems before those columns are normalized and windowed for model training. The pattern here mirrors the earlier plots of log returns and volumes, where integer-based iloc slices are reused to select contiguous feature groups for plotting, and it ties directly into the subsequent correlation analysis that reuses the same volatility column slice to compare volatility against volume. Using iloc ensures selection by column position (not name), producing a consistent, repeatable set of subplots for the volatility block.
In [9] Copy
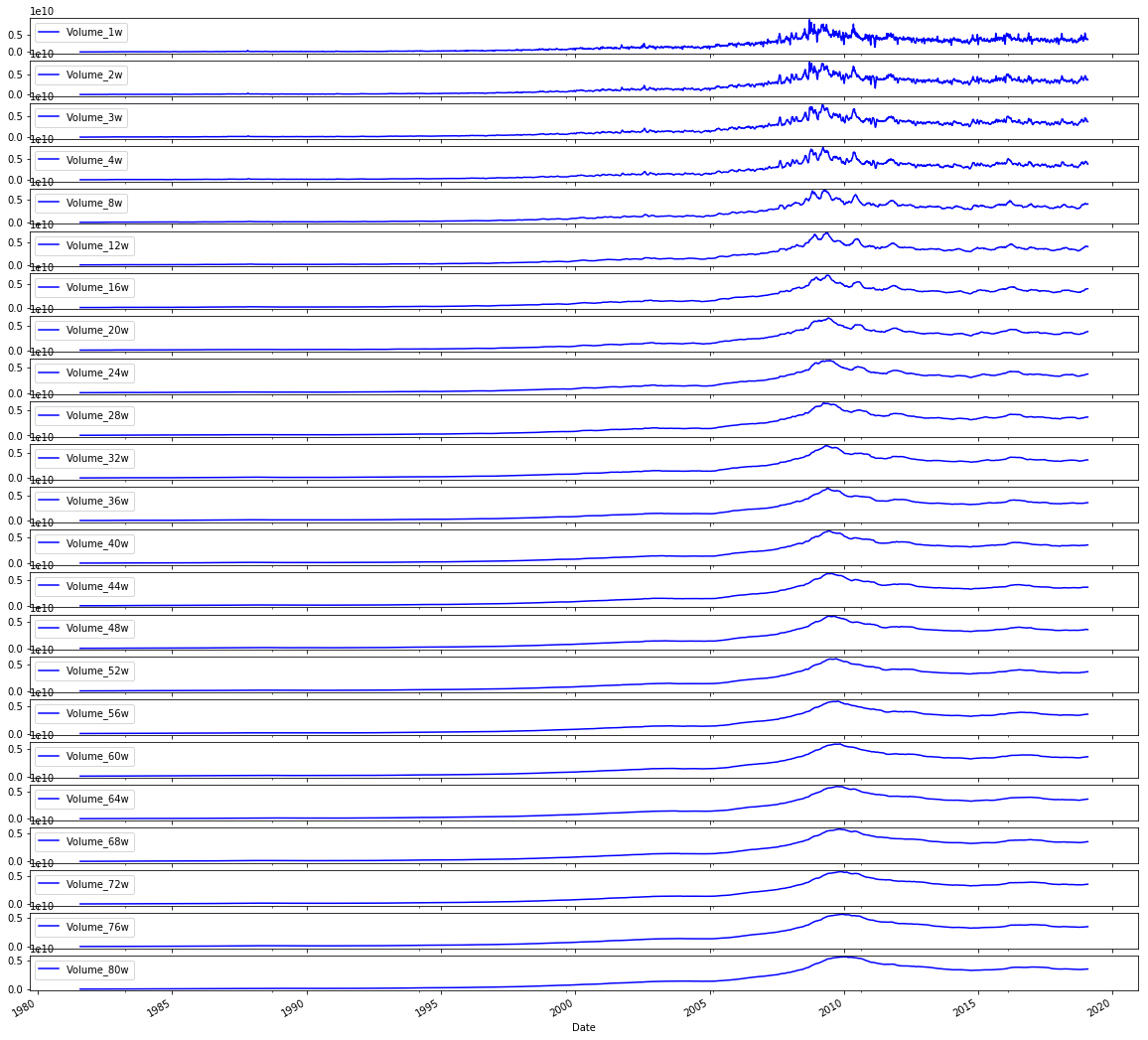
# Plot the Volumes
sp500.iloc[:,47:70].plot(subplots=True, color='blue', figsize=(20, 20));Output
This line takes the engineered sp500 DataFrame and, using positional indexing via iloc, selects the block of columns that hold the volume-derived features (the set of columns located in the column positions around the high-40s through the 60s) and then invokes the DataFrame plot method to render each of those series as its own subplot with a large figure size and a consistent blue line color. In the context of the end-to-end notebook, its role is purely exploratory: after feature engineering (for example, the Log_Ret_1d series we created earlier and the volatility features produced in other steps), it gives a visual sanity check on the temporal behavior of the volume features so you can spot trends, regime changes, outliers, or nonâstationarity before normalization, window framing, and model training. This follows the same pattern used earlier to visualize volatilities and log returns, differing only in the positional slice of columns it displays (the volatility plot used the earlier midârange column slice, and the returns plot used the initial slice), and it ties into the subsequent correlation analysis by making the volume time series explicit for inspection prior to computing volatilityâvsâvolume correlations. The use of iloc here depends on the column ordering produced by the feature engineering pipeline so that feature groups can be selected by contiguous positional ranges rather than by explicit column names.
In [10] Copy
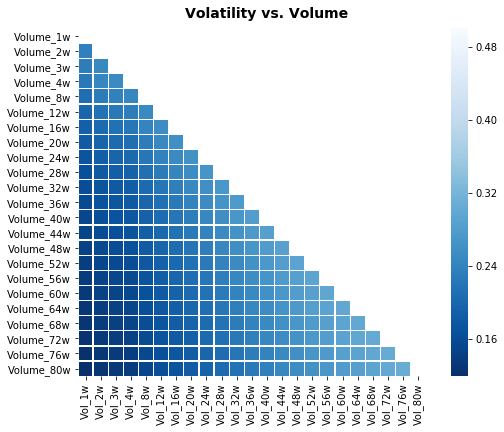
# Plot correlation matrix
focus_cols=sp500.iloc[:,24:47].columns
corr=sp500.iloc[:,24:70].corr().filter(focus_cols).drop(focus_cols)
mask=np.zeros_like(corr); mask[np.triu_indices_from(mask)]=True # we use mask to plot only part of the matrix
heat_fig, (ax)=plt.subplots(1, 1, figsize=(9,6))
heat=sns.heatmap(corr,
ax=ax,
mask=mask,
vmax=.5,
square=True,
linewidths=.2,
cmap="Blues_r")
heat_fig.subplots_adjust(top=.93)
heat_fig.suptitle('Volatility vs. Volume', fontsize=14, fontweight='bold')
plt.savefig('heat1.eps', dpi=200, format='eps');Output
The focus_cols assignment extracts the names of a contiguous block of columns from the sp500 DataFrame â specifically the columns that hold the volatility-related features the notebook wants to examine. Remember the sp500 DataFrame was fetched from Yahoo Finance earlier and that the pipeline already added engineered series such as Log_Ret_1d; focus_cols is just grabbing the column labels for the volatility block (by positional indexing) so they can be treated as the “target” variables for the correlation view. The corr computation then builds a correlation matrix over a slightly larger slice of features and selects only the columns matching focus_cols while removing the rows with those same labels, producing a rectangular table of pairwise correlations between the volatility block and the other features in the selected window (this avoids plotting self-correlations on the diagonal). A boolean mask is created from an all-zero array with its upper triangle set to true so the heatmap will visually suppress half of the symmetric correlation table. Finally, the code creates a matplotlib figure, renders the seaborn heatmap with the mask and a blue colormap, annotates the plot title to indicate this is the Volatility versus Volume comparison, and writes the figure to disk. The similar snippets elsewhere follow the same pattern â they pick a different contiguous block into focus_cols and change the source slice used for the correlation matrix (and the title), so each heatmap highlights different cross-relationships (e.g., volatility vs. return or volume vs. return); one related snippet instead plots the raw volatility time series rather than a correlation heatmap.
In [11] Copy
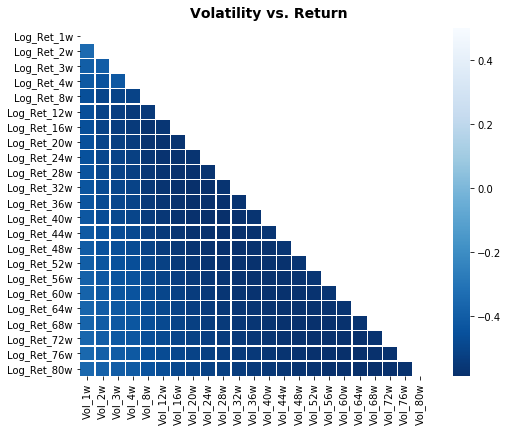
# Plot correlation matrix
focus_cols=sp500.iloc[:,24:47].columns
corr=sp500.iloc[:,1:47].corr().filter(focus_cols).drop(focus_cols)
mask=np.zeros_like(corr); mask[np.triu_indices_from(mask)]=True # we use mask to plot only part of the matrix
heat_fig, (ax)=plt.subplots(1, 1, figsize=(9,6))
heat=sns.heatmap(corr,
ax=ax,
mask=mask,
vmax=.5,
square=True,
linewidths=.2,
cmap="Blues_r")
heat_fig.subplots_adjust(top=.93)
heat_fig.suptitle('Volatility vs. Return', fontsize=14, fontweight='bold')
plt.savefig('heat2.eps', dpi=200, format='eps');Output
focus_cols is assigned by taking a positional slice out of the sp500 DataFrame via iloc and then pulling the column labels for that slice; those labels represent the block of engineered volatility features created earlier in the preprocessing stage. The notebook captures the names rather than copying the data so they can be used as a selector for downstream operations â in this case to filter the corr DataFrame and to drop the volatility columns from their own axis before drawing the heatmap, which makes the visualization show relationships between volatility features and the return-oriented columns. This pattern mirrors the other heatmap blocks in the notebook (the volume vs. return and volatility vs. volume visualizations) and differs only in which contiguous column range is selected and which additional columns are included when building the correlation matrix.
In [12] Copy
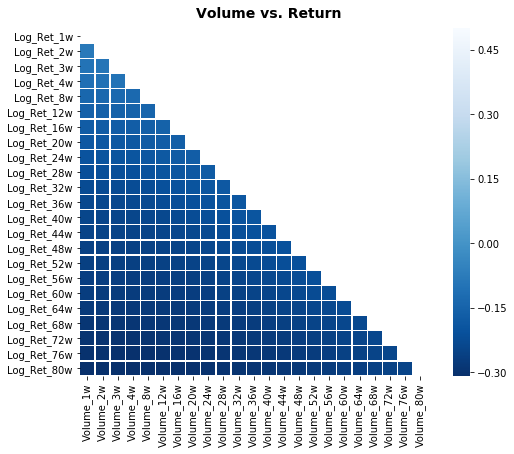
# Plot correlation matrix
focus_cols=sp500.iloc[:,47:70].columns
corr=sp500.iloc[:, np.r_[1:24, 47:70]].corr().filter(focus_cols).drop(focus_cols)
mask=np.zeros_like(corr); mask[np.triu_indices_from(mask)]=True # we use mask to plot only part of the matrix
heat_fig, (ax)=plt.subplots(1, 1, figsize=(9,6))
heat=sns.heatmap(corr,
ax=ax,
mask=mask,
vmax=.5,
square=True,
linewidths=.2,
cmap="Blues_r")
heat_fig.subplots_adjust(top=.93)
heat_fig.suptitle('Volume vs. Return', fontsize=14, fontweight='bold')
plt.savefig('heat3.eps', dpi=200, format='eps');Output
focus_cols is being set to the list of column names that represent the contiguous block of volume-derived features inside the sp500 DataFrame by using positional indexing via iloc and then reading the columns attribute. In the notebookâs workflow this variable serves as the selector for the right-hand side of the correlation analysis that follows: later code computes a correlation matrix between a set of return-related columns and other feature groups, then uses focus_cols to filter that matrix down so the heatmap only shows correlations involving the chosen volume features (and to drop the trivial self-correlations). This mirrors the same pattern used earlier when focus_cols was set to the volatility feature block, but with a different positional slice: the earlier focus_cols targeted volatility columns in the mid-20s to mid-40s, whereas here focus_cols targets the later block of positionally adjacent volume columns. In short, focus_cols isolates the names of the volume feature group from the engineered sp500 table so the plotting and correlation-filtering steps can be driven by a clear, reusable label set that supports the exploratory comparison of volume versus return signals before modeling.
4.5 Separate Test Data & Generate Model Sets for Baseline and LSTM Models
In [13] Copy
# Model Set 1: Volatility
# Baseline
X_train_1, X_test_1, y_train_1, y_test_1=train_test_split(sp500.iloc[:,24:47], sp500.iloc[:,70], test_size=0.1 ,shuffle=False, stratify=None)
# LSTM
# Input arrays should be shaped as (samples or batch, time_steps or look_back, num_features):
X_train_1_lstm=X_train_1.values.reshape(X_train_1.shape[0], 1, X_train_1.shape[1])
X_test_1_lstm=X_test_1.values.reshape(X_test_1.shape[0], 1, X_test_1.shape[1])
# Model Set 2: Return
X_train_2, X_test_2, y_train_2, y_test_2=train_test_split(sp500.iloc[:,1:24], sp500.iloc[:,70], test_size=0.1 ,shuffle=False, stratify=None)
# LSTM
# Input arrays should be shaped as (samples or batch, time_steps or look_back, num_features):
X_train_2_lstm=X_train_2.values.reshape(X_train_2.shape[0], 1, X_train_2.shape[1])
X_test_2_lstm=X_test_2.values.reshape(X_test_2.shape[0], 1, X_test_2.shape[1])
# Model Set 3: Volume
X_train_3, X_test_3, y_train_3, y_test_3=train_test_split(sp500.iloc[:,47:70], sp500.iloc[:,70], test_size=0.1 ,shuffle=False, stratify=None)
# LSTM
# Input arrays should be shaped as (samples or batch, time_steps or look_back, num_features):
X_train_3_lstm=X_train_3.values.reshape(X_train_3.shape[0], 1, X_train_3.shape[1])
X_test_3_lstm=X_test_3.values.reshape(X_test_3.shape[0], 1, X_test_3.shape[1])
# Model Set 4: Volatility and Return
X_train_4, X_test_4, y_train_4, y_test_4=train_test_split(sp500.iloc[:,1:47], sp500.iloc[:,70], test_size=0.1 ,shuffle=False, stratify=None)
# LSTM
# Input arrays should be shaped as (samples or batch, time_steps or look_back, num_features):
X_train_4_lstm=X_train_4.values.reshape(X_train_4.shape[0], 1, X_train_4.shape[1])
X_test_4_lstm=X_test_4.values.reshape(X_test_4.shape[0], 1, X_test_4.shape[1])
# Model Set 5: Volatility and Volume
X_train_5, X_test_5, y_train_5, y_test_5=train_test_split(sp500.iloc[:,24:70], sp500.iloc[:,70], test_size=0.1 ,shuffle=False, stratify=None)
# LSTM
# Input arrays should be shaped as (samples or batch, time_steps or look_back, num_features):
X_train_5_lstm=X_train_5.values.reshape(X_train_5.shape[0], 1, X_train_5.shape[1])
X_test_5_lstm=X_test_5.values.reshape(X_test_5.shape[0], 1, X_test_5.shape[1])
# Model Set 6: Return and Volume
X_train_6, X_test_6, y_train_6, y_test_6=train_test_split(pd.concat([sp500.iloc[:,1:24], sp500.iloc[:,47:70]], axis=1), sp500.iloc[:,70], test_size=0.1 ,shuffle=False, stratify=None)
# LSTM
# Input arrays should be shaped as (samples or batch, time_steps or look_back, num_features):
X_train_6_lstm=X_train_6.values.reshape(X_train_6.shape[0], 1, X_train_6.shape[1])
X_test_6_lstm=X_test_6.values.reshape(X_test_6.shape[0], 1, X_test_6.shape[1])
# Model Set 7: Volatility, Return and Volume
X_train_7, X_test_7, y_train_7, y_test_7=train_test_split(sp500.iloc[:,1:70], sp500.iloc[:,70], test_size=0.1 ,shuffle=False, stratify=None)
# LSTM
# Input arrays should be shaped as (samples or batch, time_steps or look_back, num_features):
X_train_7_lstm=X_train_7.values.reshape(X_train_7.shape[0], 1, X_train_7.shape[1])
X_test_7_lstm=X_test_7.values.reshape(X_test_7.shape[0], 1, X_test_7.shape[1])The statement uses train_test_split to carve a temporally consistent training and holdout set from the sp500 DataFrame by selecting the contiguous block of columns that hold the volatility-derived predictors (the columns positioned in the mid-20s through mid-40s) as X and a single target column (the column at position 70) as y, then reserving the final 10% of rows for testing while explicitly disabling shuffling and not applying stratification. Conceptually, this moves the pipeline from feature engineering (where you computed log returns and multi-week volatility measures such as Vol_16w) into the modeling stage by producing the four arrays X_train_1, X_test_1, y_train_1 and y_test_1 that downstream code will use to fit a baseline learner; the LSTM branch later consumes the same split but reshapes the X arrays into the threeâdimensional form an LSTM expects. The same temporal split pattern is then repeated for other model sets (returns, volume, and their combinations), differing only in which column slices are chosen for X (including one case that concatenates two noncontiguous slices), so this call implements the canonical, time-aware train/test partition for the volatility-focused experiment.
4.6 Show Label Distribution
In [14] Copy
print("train set increase bias = "+ str(np.mean(y_train_7==1))+"%")
print("test set increase bias = " + str(np.mean(y_test_7==1))+"%")Output
[stdout]
train set increase bias = 0.6243535496003761%
test set increase bias = 0.6437632135306554%These two lines print a simple labelâdistribution check for the sevenâday horizon labels so you can see how many examples are labeled as an “increase” in the training and test partitions before any models are trained. Concretely, the notebook computes the proportion of entries in y_train_7 that are equal to the positive class (using numpyâs mean over a boolean indicator) and reports that as the training set âincreaseâ bias, then does the same for y_test_7 to report the test set bias. In the endâtoâend experimental pipeline this is a lightweight, early diagnostic: it reveals class imbalance and any shift in the positiveâlabel prior between the temporally split train and test sets, information that directly affects how you interpret downstream results from LSTM and classical learners. This differs from the later blocks youâve seen (the confusion matrix + accuracy/precision/recall block that calls tuned_model_7_b.predict and the ROC block that uses tuned_model_7_b.predict_proba) because those evaluate model performance using predictions versus true labels, whereas these prints only summarize the trueâlabel priors themselves.
5. Workflow

Walk Forward Cross-Validation
Time Series cross-validator provides train/dev indices to split time series data samples that are observed at fixed time intervals, in train/dev sets. In each split, dev indices must be higher than before, and thus shuffling in cross validator is inappropriate. The following graph illustrates how the time series split works:

In [15] Copy
# Time Series Split
dev_size=0.1
n_splits=int((1//dev_size)-1) # using // for integer division
tscv=TimeSeriesSplit(n_splits=n_splits)dev_size is the fraction of the whole time series reserved for each validation (dev) fold when performing walkâforward crossâvalidation; setting dev_size to 0.1 means each dev fold will be roughly 10% of the sample horizon. The notebook turns that proportion into the number of sequential splits by taking the integer quotient of one divided by dev_size and subtracting one, which with 0.1 yields eight TimeSeriesSplit folds, so TimeSeriesSplit is constructed with eight splits. TimeSeriesSplit then produces temporally ordered train/dev pairs where each dev block sits later in time than the previous ones (shuffling is therefore inappropriate), implementing the walkâforward scheme illustrated in the diagram. In practice that means all of the engineered features you prepared earlier (for example the multiweek cumulative log returns like Log_Ret_8w and Log_Ret_20w and the rolling volatility Vol_16w) will be evaluated in a temporally consistent way: for each of the eight folds the preprocessing Pipeline (for example the StandardScaler inside pipeline_b) and any hyperparameter search will be refit only on the subâtraining portion and validated on the subsequent 10% dev window, so no future information from the dev window leaks into training or scaling. dev_size therefore controls the temporal granularity of the outer crossâvalidation loop (unlike parameters such as num_samples or look_back, which control model input shapes), and by choosing 0.1 the notebook creates a modest number of progressive train/dev rounds to compare LSTM and classical learners across time.
Scaling (Standardization/Normalization)
The splitting of the data set during cross-validation should be done before doing any preprocessing. Any process that extracts knowledge from the dataset should only ever be applied to the training portion of the data set, so any cross-validation should be the âoutermost loopâ in our processing.
The Pipeline class is a class that allows âgluingâ together multiple processing steps into a single scikit-learn estimator. The Pipeline class itself has fit, predict and score methods and behaves just like any other model in scikit-learn. The most common use-case of the pipeline class is in chaining preprocessing steps (like scaling of the data) together with a supervised model like a classifier.
For each split in the Walk-Forward CV, the Scaler is refit with only the Sub training splits, not leaking any information of the test split into the parameter search, as illustrated below.

The preprocessing includes the following steps:
Step 0: The data is split into TRAINING data and VALIDATION data according to the cv parameter that we specified in the GridSearchCV or RandomizedSearchCV
Step 1: the scaler is fitted on the TRAINING data
Step 2: the scaler transforms TRAINING data
Step 3: the models are fitted/trained using the transformed TRAINING data
Step 4: the scaler is used to transform the VALIDATION data
Step 5: the trained models predict using the transformed VALIDATION data
Step 6: the scaler is fitted on the TRAINING and VALIDATION data
Step 7: the scaler transforms TRAINING and VALIDATION data
Step 8: the model is fitted/trained using the transformed TRAINING and VALIDATION data and the best found parameters during Walk-Forward CV
Step 9: the scaler transforms TEST data
Step 10: the trained model predicts using the transformed TEST data
6 Regularization
Regularization adds a penalty on the different parameters of the baseline model to reduce the freedom of the model. Hence, the model will be less likely to fit the noise of the training data and will improve the generalization abilities of the mode
6.1 L2 Regularization (Ridge penalisation)
The L2 regularization adds a penalty equal to the sum of the squared value of the coefficients.
J=1m∑i=1mL(ai,yi)+λ∑i=1mβi2(1) J = \frac{1}{m} \sum_{i=1}^m \mathcal{L}(a_{i}, y_{i}) + \lambda\sum_{i=1}^m\beta_{i}^2\tag{1} J=m1i=1∑mL(ai,yi)+λi=1∑mβi2(1)
The λ parameter is a scalar that should be learned as well, using walk-forward-cross-validation.
L2 regularization will force the parameters to be relatively small, the bigger the penalization, the smaller (and the more robust) the coefficients are.

6.2 L1 Regularization (Lasso penalisation)
The L1 regularization adds a penalty equal to the sum of the absolute value of the coefficients.
J=1m∑i=1mL(ai,yi)+λ∑i=1m∣βi∣(2) J = \frac{1}{m} \sum_{i=1}^m \mathcal{L}(a_{i}, y_{i}) + \lambda\sum_{i=1}^m|\beta_{i}|\tag{2} J=m1i=1∑mL(ai,yi)+λi=1∑m∣βi∣(2)
The λ parameter is a scalar that should be learned as well, using walk-forward-cross-validation.
L1 regularization will shrink some parameters to zero. Hence some variables will not play any role in the model, L1 regression can be seen as a way to select features in a model.

6.3 Elastic Net
Elastic-net is a mix of both L1 and L2 regularizations. A penalty is applied to the sum of the absolute values and to the sum of the squared values:
J=1m∑i=1mL(ai,yi)+λ((1−α)∑i=1mβi2+α∑i=1m∣βi∣)(3) J = \frac{1}{m} \sum_{i=1}^m \mathcal{L}(a_{i}, y_{i}) + \lambda((1-\alpha)\sum_{i=1}^m\beta{i}^2+\alpha\sum_{i=1}^m|\beta_{i}|)\tag{3} J=m1i=1∑mL(ai,yi)+λ((1−α)i=1∑mβi2+αi=1∑m∣βi∣)(3)
Lambda is a shared penalization parameter while alpha sets the ratio between L1 and L2 regularization in the Elastic Net Regularization. Hence, we expect a hybrid behavior between L1 and L2 regularization.
Though coefficients are cut, the cut is less abrupt than the cut with lasso penalization alone.
Models
Configuration Baseline Models
In [16] Copy
# Standardized Data
steps_b=[('scaler', StandardScaler(copy=True, with_mean=True, with_std=True)),
('logistic', linear_model.SGDClassifier(loss="log", shuffle=False, early_stopping=False, tol=1e-3, random_state=1))]
#Normalized Data
#steps_b=[('scaler', MinMaxScaler(feature_range=(0, 1), copy=True)),
# ('logistic', linear_model.SGDClassifier(loss="log", shuffle=False, early_stopping=False, tol=1e-3, random_state=1))]
pipeline_b=Pipeline(steps_b) # Using a pipeline we glue together the Scaler & the Classifier
# This ensure that during cross validation the Scaler is fitted to only the training folds
# Penalties
penalty_b=['l1', 'l2', 'elasticnet']
# Evaluation Metric
scoring_b={'AUC': 'roc_auc', 'accuracy': make_scorer(accuracy_score)} #multiple evaluation metrics
metric_b='accuracy' #scorer is used to find the best parameters for refitting the estimator at the endThis section wires a preprocessing-and-estimator chain that the notebook reuses during the model selection and evaluation phase so that classical learners are compared fairly against the LSTM on time-windowed examples. It first picks a StandardScaler configured to center each feature and scale to unit variance so features like log-returns and lagged observations are on a common numerical scale (an important step because the downstream learner is an SGD-based logistic classifier which is sensitive to feature scales). An alternative MinMaxScaler is kept commented out for experiments that prefer a 0â1 normalization instead of z-score standardization. The scaler is then glued to a linear_model.SGDClassifier that optimizes a logistic loss; placing the scaler before the classifier inside a sklearn Pipeline guarantees that, during walk-forward cross-validation, the scaling parameters are always estimated only on the training folds and never leak information from the validation/test folds. penalty_b is defined as the set of regularization priors that will be swept later during hyperparameter search, and scoring_b collects both ROC AUC and an accuracy scorer so GridSearchCV can evaluate multiple metrics while metric_b designates accuracy as the single metric used to pick the final refit model. pipeline_b is therefore the reusable estimator object that the subsequent GridSearchCV/RandomizedSearchCV blocks (the ones you saw in the iterations_1_b and iterations_2_b pattern) accept as their estimator argument, enabling consistent, leak-free preprocessing inside each time-series CV fold.
Configuration LSTM Models
In [17] Copy
# Batch_input_shape=[1, 1, Z] -> (batch size, time steps, number of features)
# Data set inputs(trainX)=[X, 1, Z] -> (samples, time steps, number of features)
# number of samples
num_samples=1
# time_steps
look_back=1
# Evaluation Metric
scoring_lstm='accuracy'Within the endâtoâend notebook’s LSTM configuration, num
6. Models
Model 1: Volatility
Baseline
In [18] Copy
# Model specific Parameter
# Number of iterations
iterations_1_b=[8]
# Grid Search
# Regularization
alpha_g_1_b=[0.0011, 0.0013, 0.0014] #0.0011, 0.0012, 0.0013
l1_ratio_g_1_b=[0, 0.2, 0.4, 0.6, 0.8, 1]
# Create hyperparameter options
hyperparameters_g_1_b={'logistic__alpha':alpha_g_1_b,
'logistic__l1_ratio':l1_ratio_g_1_b,
'logistic__penalty':penalty_b,
'logistic__max_iter':iterations_1_b}
# Create grid search
search_g_1_b=GridSearchCV(estimator=pipeline_b,
param_grid=hyperparameters_g_1_b,
cv=tscv,
verbose=0,
n_jobs=-1,
scoring=scoring_b,
refit=metric_b,
return_train_score=False)
# Setting refit='Accuracy', refits an estimator on the whole dataset with the parameter setting that has the best cross-validated mean Accuracy score.
# For multiple metric evaluation, this needs to be a string denoting the scorer is used to find the best parameters for refitting the estimator at the end
# If return_train_score=True training results of CV will be saved as well
# Fit grid search
tuned_model_1_b=search_g_1_b.fit(X_train_1, y_train_1)
#search_g_1_b.cv_results_
# Random Search
# Create regularization hyperparameter distribution using uniform distribution
#alpha_r_1_b=uniform(loc=0.00006, scale=0.002)
#l1_ratio_r_1_b=uniform(loc=0, scale=1)
# Create hyperparameter options
#hyperparameters_r_1_b={'logistic__alpha':alpha_r_1_b, 'logistic__l1_ratio':l1_ratio_r_1_b, 'logistic__penalty':penalty_b,'logistic__max_iter':iterations_1_b}
# Create randomized search
#search_r_1_b=RandomizedSearchCV(pipeline_b,
# hyperparameters_r_1_b,
# n_iter=10,
# random_state=1,
# cv=tscv,
# verbose=0,
# n_jobs=-1,
# scoring=scoring_b,
# refit=metric_b,
# return_train_score=True)
# Setting refit='Accuracy', refits an estimator on the whole dataset with the parameter setting that has the best cross-validated Accuracy score.
# For multiple metric evaluation, this needs to be a string denoting the scorer is used to find the best parameters for refitting the estimator at the end
# If return_train_score=True training results of CV will be saved as well
# Fit randomized search
#tuned_model_1_b=search_r_1_b.fit(X_train_1, y_train_1)
# View Cost function
print('Loss function:', tuned_model_1_b.best_estimator_.get_params()['logistic__loss'])
# View Accuracy
print(metric_b +' of the best model: ', tuned_model_1_b.best_score_);print("\n")
# best_score_ Mean cross-validated score of the best_estimator
# View best hyperparameters
print("Best hyperparameters:")
print('Number of iterations:', tuned_model_1_b.best_estimator_.get_params()['logistic__max_iter'])
print('Penalty:', tuned_model_1_b.best_estimator_.get_params()['logistic__penalty'])
print('Alpha:', tuned_model_1_b.best_estimator_.get_params()['logistic__alpha'])
print('l1_ratio:', tuned_model_1_b.best_estimator_.get_params()['logistic__l1_ratio'])
# Find the number of nonzero coefficients (selected features)
print("Total number of features:", len(tuned_model_1_b.best_estimator_.steps[1][1].coef_[0][:]))
print("Number of selected features:", np.count_nonzero(tuned_model_1_b.best_estimator_.steps[1][1].coef_[0][:]))
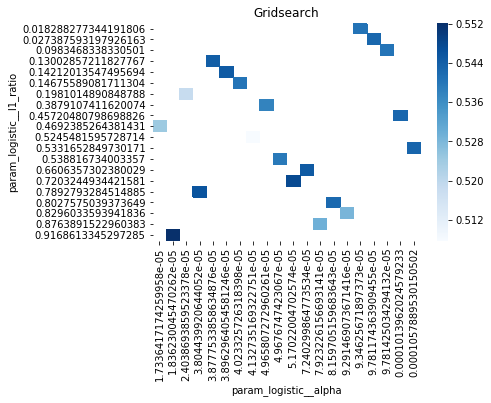
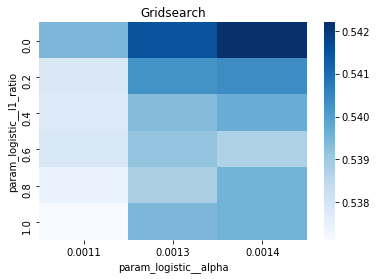
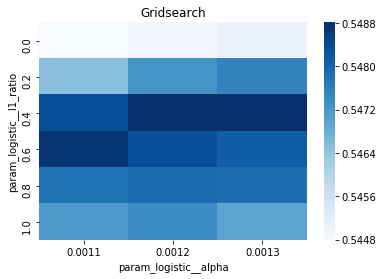
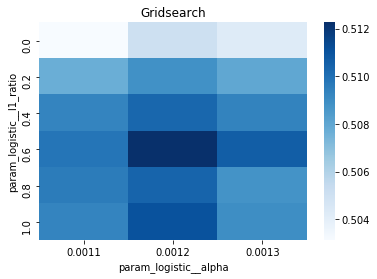
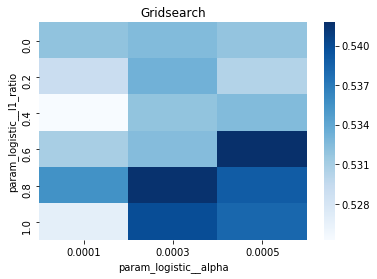
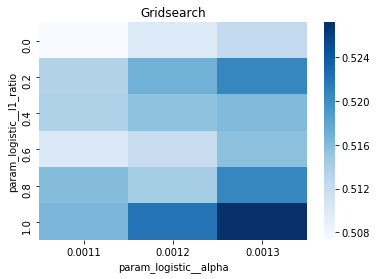
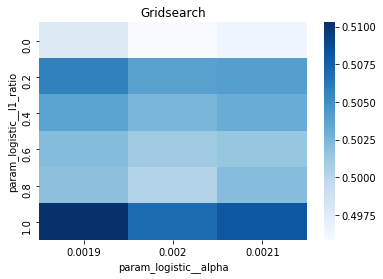
# Gridsearch table
plt.title('Gridsearch')
pvt_1_b=pd.pivot_table(pd.DataFrame(tuned_model_1_b.cv_results_), values='mean_test_accuracy', index='param_logistic__l1_ratio', columns='param_logistic__alpha')
ax_1_b=sns.heatmap(pvt_1_b, cmap="Blues")
plt.show()Output
[stdout]
Loss function: log
accuracy of the best model: 0.5448412698412698
Best hyperparameters:
Number of iterations: 8
Penalty: l2
Alpha: 0.0014
l1_ratio: 0
Total number of features: 23
Number of selected features: 23For the endâtoâend experiment in this notebook, iterations_1_b is the modelâspecific declaration of the maximumâiteration budget that the logistic component of pipeline_b will be allowed during hyperparameter search. The notebook assembles a hyperparameter dictionary that includes the logistic maxâiteration setting alongside regularization candidates (alpha and l1_ratio) and the penalty, and then hands that dictionary to GridSearchCV together with the timeâseries crossâvalidator tscv, the scoring collection scoring_b, and the refit metric metric_b. Because iterations_1_b is provided as a oneâelement list containing the value eight, the grid search will only consider a maxâiteration value of eight for the logistic solver when it evaluates every combination of the other regularization parameters; the fitted search object tuned_model_1_b therefore records the best combination found under that iteration budget and exposes it via best_estimator_.get_params() and best_score_. This follows the same pattern used elsewhere in the notebook: each model variant defines its own iterations_N_b list and matching regularization ranges, constructs a hyperparameters mapping, runs GridSearchCV (with a commented RandomizedSearchCV alternative available), fits against its modelâspecific training split, and then inspects convergence and selection statistics. The difference between iterations_1_b and the analogous variables for other models is simply the numeric budget chosen (for example, some models use ten), reflecting that each model block sets its own solver iteration parameter before running the same crossâvalidated search-andârefit flow.
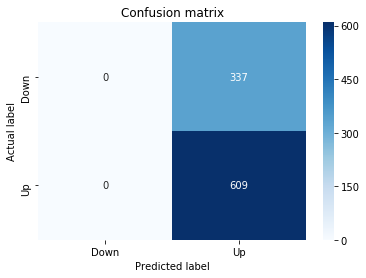
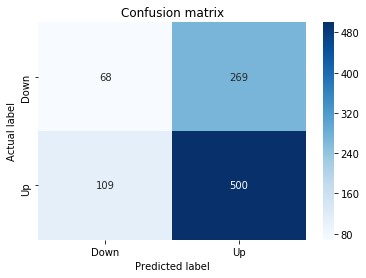
Confusion Matrix
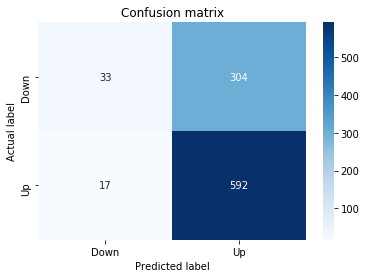
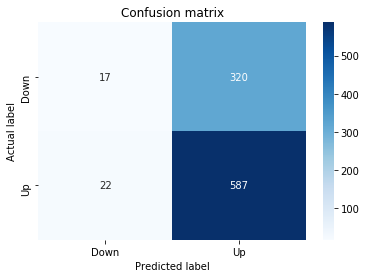
In [19] Copy
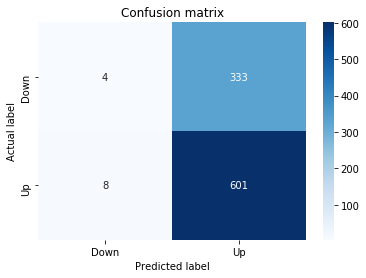
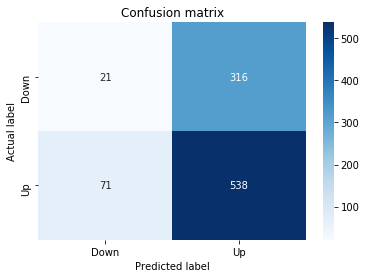
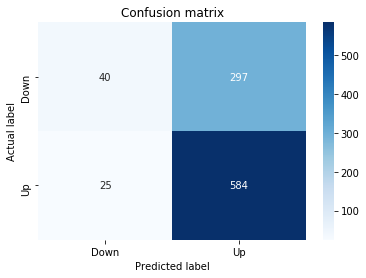
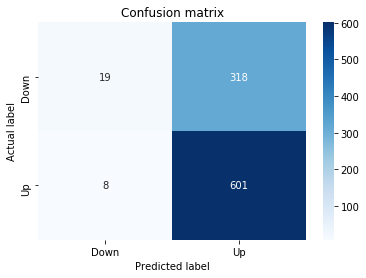
# Make predictions
y_pred_1_b=tuned_model_1_b.predict(X_test_1)
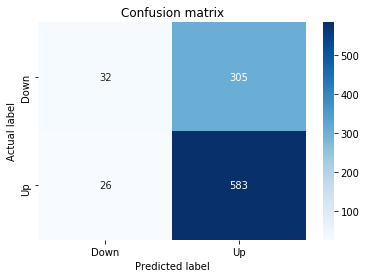
# create confustion matrix
fig, ax=plt.subplots()
sns.heatmap(pd.DataFrame(metrics.confusion_matrix(y_test_1, y_pred_1_b)), annot=True, cmap="Blues" ,fmt='g')
plt.title('Confusion matrix'); plt.ylabel('Actual label'); plt.xlabel('Predicted label')
ax.xaxis.set_ticklabels(['Down', 'Up']); ax.yaxis.set_ticklabels(['Down', 'Up'])
print("Accuracy:",metrics.accuracy_score(y_test_1, y_pred_1_b))
print("Precision:",metrics.precision_score(y_test_1, y_pred_1_b))
print("Recall:",metrics.recall_score(y_test_1, y_pred_1_b))Output
[stdout]
Accuracy: 0.6501057082452432
Precision: 0.6565315315315315
Recall: 0.9573070607553367To evaluate the first tuned estimator on heldâout data, the notebook invokes the predictor associated with tuned_model_1_b to produce a sequence of predicted class labels for the preprocessed test examples contained in X_test_1, and those predictions are captured as y_pred_1_b. tuned_model_1_b is the hyperparameterâtuned learner trained earlier in the pipeline (either an LSTM wrapper or a classical classifier depending on the experiment), and X_test_1 consists of the timeâwindowed, normalized feature sequences assembled from the engineered return series (for example the features you already saw listed such as Log_Ret_4w, Log_Ret_20w and Log_Ret_40w in the DataFrame info outputs). The output y_pred_1_b is therefore the modelâs binary Up/Down forecast for each test sequence over the chosen horizon; those predicted labels are immediately consumed by the subsequent confusionâmatrix plotting and the accuracy/precision/recall calculations. This call is the same operational pattern used for tuned_model_2_b, tuned_model_3_b and tuned_model_4_b, differing only in which tuned estimator and corresponding X_test partition are being evaluated.
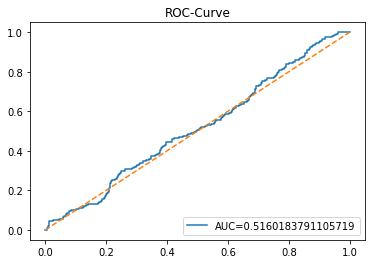
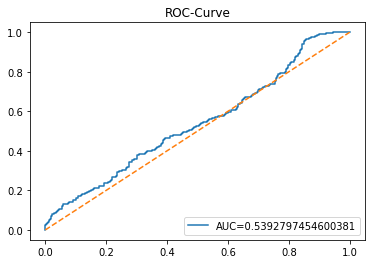
ROC Curve
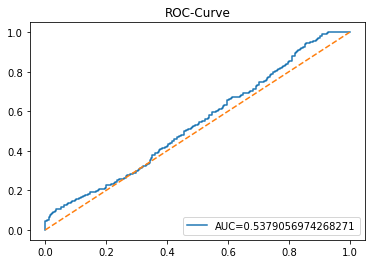
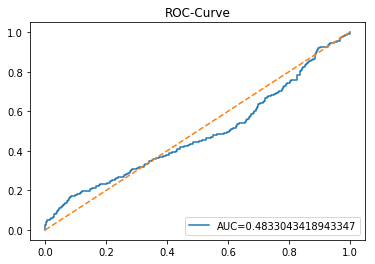
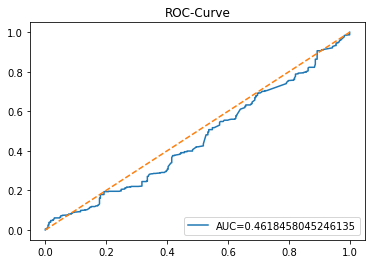
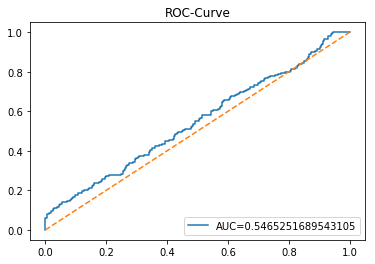
In [20] Copy
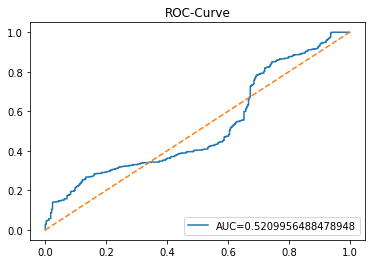
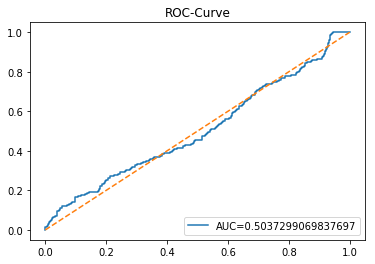
y_proba_1_b=tuned_model_1_b.predict_proba(X_test_1)[:, 1]
fpr, tpr, _=metrics.roc_curve(y_test_1, y_proba_1_b)
auc=metrics.roc_auc_score(y_test_1, y_proba_1_b)
plt.plot(fpr,tpr,label="AUC="+str(auc))
plt.legend(loc=4)
plt.plot([0, 1], [0, 1], linestyle='--') # plot no skill
plt.title('ROC-Curve')
plt.show()Output
Here we take the previously trained and hyperparameterâtuned classifier named tuned_model_1_b and ask it to produce probability estimates for every example in the test partition X_test_1 (the temporally heldâout, windowed feature sequences created earlier in the preprocessing stage). The model’s probability output gives, for each test row, a probability distribution over the two label classes; we extract the probability assigned to the positive or “increase” class because the ROC computations that follow require continuous scores rather than hard class labels. That vector of positiveâclass probabilities is what flows into metrics.roc_curve and metrics.roc_auc_score to build the ROC curve and compute area under curve for the forecasting horizon represented by the 1 split. This call mirrors the pattern used for tuned_model_2_b and tuned_model_7_b elsewhere in the notebook, where each tuned model is asked for its positiveâclass probabilities on its corresponding X_test partition so we can compare discrimination performance across horizons.
LSTM Model
In [21] Copy
start=time.time()
# number of epochs
epochs=1
# number of units
LSTM_units_1_lstm=195
# numer of features
num_features_1_lstm=X_train_1.shape[1]
# Regularization
dropout_rate=0.1
recurrent_dropout=0.1 # 0.21
# print
verbose=0
#hyperparameter
batch_size=[1]
# hyperparameter
hyperparameter_1_lstm={'batch_size':batch_size}
# create Classifier
clf_1_lstm=KerasClassifier(build_fn=create_shallow_LSTM,
epochs=epochs,
LSTM_units=LSTM_units_1_lstm,
num_samples=num_samples,
look_back=look_back,
num_features=num_features_1_lstm,
dropout_rate=dropout_rate,
recurrent_dropout=recurrent_dropout,
verbose=verbose)
# Gridsearch
search_1_lstm=GridSearchCV(estimator=clf_1_lstm,
param_grid=hyperparameter_1_lstm,
n_jobs=-1,
cv=tscv,
scoring=scoring_lstm, # accuracy
refit=True,
return_train_score=False)
# Fit model
tuned_model_1_lstm=search_1_lstm.fit(X_train_1_lstm, y_train_1, shuffle=False, callbacks=[reset])
print("\n")
# View Accuracy
print(scoring_lstm +' of the best model: ', tuned_model_1_lstm.best_score_)
# best_score_ Mean cross-validated score of the best_estimator
print("\n")
# View best hyperparameters
print("Best hyperparameters:")
print('epochs:', tuned_model_1_lstm.best_estimator_.get_params()['epochs'])
print('batch_size:', tuned_model_1_lstm.best_estimator_.get_params()['batch_size'])
print('dropout_rate:', tuned_model_1_lstm.best_estimator_.get_params()['dropout_rate'])
print('recurrent_dropout:', tuned_model_1_lstm.best_estimator_.get_params()['recurrent_dropout'])
end=time.time()
print("\n")
print("Running Time:", end - start)Output
[stdout]
accuracy of the best model: 0.6101851851851852
Best hyperparameters:
epochs: 1
batch_size: 1
dropout_rate: 0.1
recurrent_dropout: 0.1
Running Time: 241.70626878738403At the top of the LSTM training block the notebook records a timestamp into the variable named start by invoking the system clock; that timestamp marks the moment immediately before the expensive work of model construction, GridSearchCV orchestration and the Keras training loop begins. Because this notebook is an end-to-end experimental pipeline that repeatedly builds, tunes and fits different LSTM variants using KerasClassifier and GridSearchCV with time-series crossâvalidation, capturing start at the outset lets the code later compute a single wallâclock duration (by subtracting start from a corresponding end timestamp recorded after fit completes) and print a Running Time for that experiment. In practical terms the start timestamp therefore brackets the portion of the dataflow where X_train is reshaped for LSTM input, create_shallow_LSTM is instantiated via KerasClassifier, and search.fit executes crossâvalidated training with callbacks; the duration is used to compare compute cost across model variants and to report how long each hyperparameter search took. You see the exact same timing pattern repeated across the other LSTM blocks in the notebook; those repetitions differ only in model hyperparameters and which X_train/X_test split they operate on, while the timing call itself is used identically to produce a comparable runtime metric for each experiment.
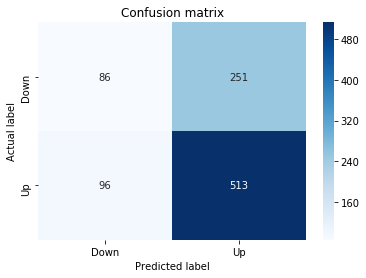
Confusion Matrix
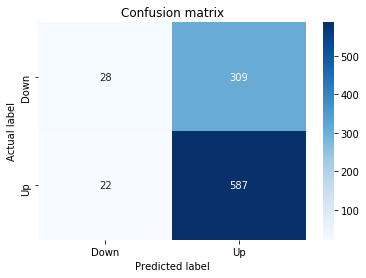
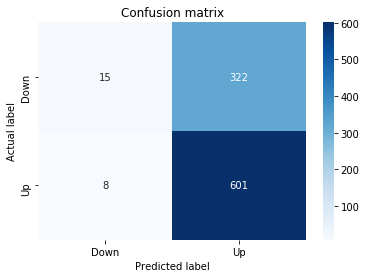
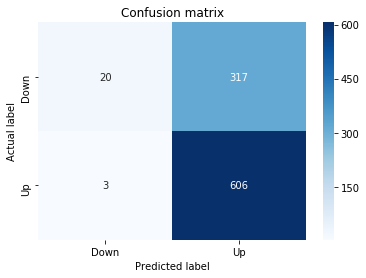
In [22] Copy
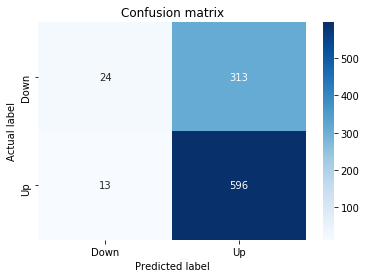
# Make predictions
y_pred_1_lstm=tuned_model_1_lstm.predict(X_test_1_lstm)
# create confustion matrix
fig, ax=plt.subplots()
sns.heatmap(pd.DataFrame(metrics.confusion_matrix(y_test_1, y_pred_1_lstm)), annot=True, cmap="Blues" ,fmt='g')
plt.title('Confusion matrix'); plt.ylabel('Actual label'); plt.xlabel('Predicted label')
ax.xaxis.set_ticklabels(['Down', 'Up']); ax.yaxis.set_ticklabels(['Down', 'Up'])
print("Accuracy:",metrics.accuracy_score(y_test_1, y_pred_1_lstm))
print("Precision:",metrics.precision_score(y_test_1, y_pred_1_lstm))
print("Recall:",metrics.recall_score(y_test_1, y_pred_1_lstm))Output
[stdout]
Accuracy: 0.6437632135306554
Precision: 0.6437632135306554
Recall: 1.0For experiment 1 the notebook asks the trained LSTM estimator tuned_model_1_lstm to produce discrete direction predictions for the temporally heldâout, windowed test set X_test_1_lstm â similar in role to the earlier classical prediction we inspected when tuned_model_1_b produced y_pred_1_b, but here the inputs are LSTM-shaped sequences created by the preprocessing stage. Those predicted labels, y_pred_1_lstm, are then compared against the true labels y_test_1 to build a confusion matrix via the metrics module; that contingency table is visualized as a blue heatmap using seaborn with the plot axes relabeled to the two target classes (Down and Up) so you can quickly see true/false positives and negatives. Finally the code prints three standard classification summaries â accuracy, precision, and recall computed from y_test_1 and y_pred_1_lstm â so the LSTMâs binary directional forecasting performance for this experimental split can be directly compared to the classical learners and to the other LSTM runs (y_pred_2_lstm, y_pred_3_lstm, y_pred_4_lstm), which follow the same predict â confusion matrix â metric printout pattern for their respective test partitions.
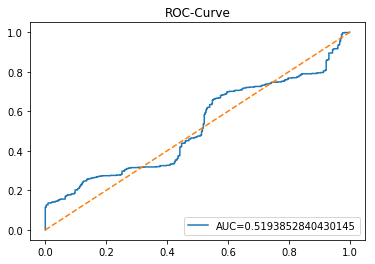
ROC Curve
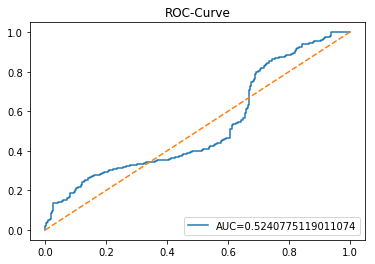
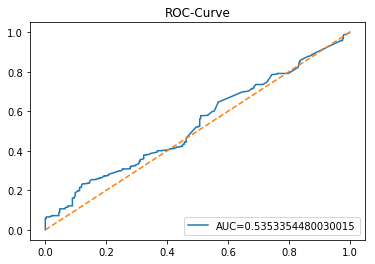
In [23] Copy
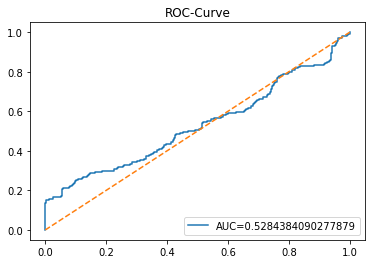
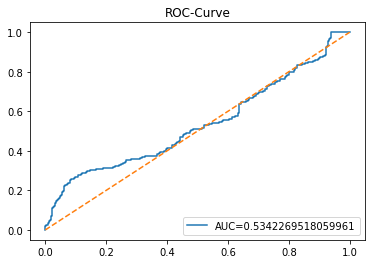
y_proba_1_lstm=tuned_model_1_lstm.predict_proba(X_test_1_lstm)[:, 1]
fpr, tpr, _=metrics.roc_curve(y_test_1, y_proba_1_lstm)
auc=metrics.roc_auc_score(y_test_1, y_proba_1_lstm)
plt.plot(fpr,tpr,label="AUC="+str(auc))
plt.legend(loc=4)
plt.plot([0, 1], [0, 1], linestyle='--') # plot no skill
plt.title('ROC-Curve')
plt.show()Output
This cell evaluates the tuned LSTM classifier tuned_model_1_lstm on the temporally heldâout, windowed test set X_test_1_lstm by asking the model for class probability estimates and then plotting an ROC curve to quantify discrimination ability. Concretely, the notebook extracts the modelâs estimated probability for the positive class for every windowed test example produced during the preprocessing stage, then feeds those probabilities and the true labels y_test_1 into sklearn.metrics.roc_curve to produce the false positive and true positive rate arrays and into sklearn.metrics.roc_auc_score to compute the scalar AUC summary. The plot call draws the ROC line labeled with the AUC, adds a legend in the lowerâright, overlays the diagonal ânoâskillâ reference, titles the figure âROCâCurveâ, and displays it. This follows the same evaluation pattern used earlier for the classical estimator tuned_model_1_b (where probabilities were computed over flat test features) and for the other tuned_model_*_lstm runs; the key difference here is that the inputs are LSTMâshaped, timeâwindowed sequences rather than tabular feature rows, so the plotted ROC measures how well the recurrent modelâs positiveâclass probabilities separate future up/down outcomes on the heldâout temporal partition.
Model 2: Return
Baseline
In [24] Copy
# Model specific Parameter
# Number of iterations
iterations_2_b=[8]
# Grid Search
# Regularization
alpha_g_2_b=[0.0011, 0.0012, 0.0013]
l1_ratio_g_2_b=[0, 0.2, 0.4, 0.6, 0.8, 1]
# Create hyperparameter options
hyperparameters_g_2_b={'logistic__alpha':alpha_g_2_b,
'logistic__l1_ratio':l1_ratio_g_2_b,
'logistic__penalty':penalty_b,
'logistic__max_iter':iterations_2_b}
# Create grid search
search_g_2_b=GridSearchCV(estimator=pipeline_b,
param_grid=hyperparameters_g_2_b,
cv=tscv,
verbose=0,
n_jobs=-1,
scoring=scoring_b,
refit=metric_b,
return_train_score=False)
# Setting refit='Accuracy', refits an estimator on the whole dataset with the parameter setting that has the best cross-validated mean Accuracy score.
# For multiple metric evaluation, this needs to be a string denoting the scorer is used to find the best parameters for refitting the estimator at the end
# If return_train_score=True training results of CV will be saved as well
# Fit grid search
tuned_model_2_b=search_g_2_b.fit(X_train_2, y_train_2)
#search_g_2_b.cv_results_
# Random Search
# Create regularization hyperparameter distribution using uniform distribution
#alpha_r_2_b=uniform(loc=0.00006, scale=0.002) #loc=0.00006, scale=0.002
#l1_ratio_r_2_b=uniform(loc=0, scale=1)
# Create hyperparameter options
#hyperparameters_r_2_b={'logistic__alpha':alpha_r_2_b, 'logistic__l1_ratio':l1_ratio_r_2_b, 'logistic__penalty':penalty_b,'logistic__max_iter':iterations_2_b}
# Create randomized search
#search_r_2_b=RandomizedSearchCV(pipeline_b, hyperparameters_r_2_b, n_iter=10, random_state=1, cv=tscv, verbose=0, n_jobs=-1, scoring=scoring_b, refit=metric_b, return_train_score=False)
# Setting refit='Accuracy', refits an estimator on the whole dataset with the parameter setting that has the best cross-validated Accuracy score.
# Fit randomized search
#tuned_model_2_b=search_r_2_b.fit(X_train_2, y_train_2)
# View Cost function
print('Loss function:', tuned_model_2_b.best_estimator_.get_params()['logistic__loss'])
# View Accuracy
print(metric_b +' of the best model: ', tuned_model_2_b.best_score_);print("\n")
# best_score_ Mean cross-validated score of the best_estimator
# View best hyperparameters
print("Best hyperparameters:")
print('Number of iterations:', tuned_model_2_b.best_estimator_.get_params()['logistic__max_iter'])
print('Penalty:', tuned_model_2_b.best_estimator_.get_params()['logistic__penalty'])
print('Alpha:', tuned_model_2_b.best_estimator_.get_params()['logistic__alpha'])
print('l1_ratio:', tuned_model_2_b.best_estimator_.get_params()['logistic__l1_ratio'])
# Find the number of nonzero coefficients (selected features)
print("Total number of features:", len(tuned_model_2_b.best_estimator_.steps[1][1].coef_[0][:]))
print("Number of selected features:", np.count_nonzero(tuned_model_2_b.best_estimator_.steps[1][1].coef_[0][:]))
# Gridsearch table
plt.title('Gridsearch')
pvt_2_b=pd.pivot_table(pd.DataFrame(tuned_model_2_b.cv_results_), values='mean_test_accuracy', index='param_logistic__l1_ratio', columns='param_logistic__alpha')
ax_2_b=sns.heatmap(pvt_2_b, cmap="Blues")
plt.show()Output
[stdout]
Loss function: log
accuracy of the best model: 0.5543650793650794
Best hyperparameters:
Number of iterations: 8
Penalty: elasticnet
Alpha: 0.0013
l1_ratio: 0.4
Total number of features: 23
Number of selected features: 10iterations_2_b is the small list that supplies the logistic solverâs maximum-iteration setting used by the Model 2 experimental workflow; in this notebook the list contains a single value of eight, so the hyperparameter grid will only try that iteration budget when GridSearchCV explores regularization choices for the pipeline named pipeline_b. Conceptually this cell takes the Model 2 training partition X_train_2 and y_train_2 and performs a time-series-aware hyperparameter search (GridSearchCV with the tscv splitter) over a small grid of alpha values and l1_ratio values together with the fixed penalty choices already defined for pipeline_b, using scoring_b to evaluate folds and refitting the final estimator according to metric_b; the fitted search object is saved as tuned_model_2_b. A commented RandomizedSearchCV alternative shows an option to sample continuous regularization distributions instead of enumerating them, but the active path uses the exhaustive grid. After fit, the code queries tuned_model_2_b.best_estimator_ to report the learned loss, the cross-validated best_score_, the selected max-iteration, penalty, alpha and l1_ratio, and then inspects the logistic stepâs coefficient vector to report total and nonzero feature counts (showing how many inputs survived regularized selection). Finally the cross-validation results are converted into a pivot table of mean test accuracy across l1_ratio by alpha and visualized with a seaborn heatmap so you can see how accuracy varied over the regularization grid. iterations_2_b=[8] is consistent with the same iteration setting used for Model 1 and Model 4 (both use eight) whereas Model 5 uses ten, which reflects a deliberate, per-model choice of solver iteration budget while keeping the rest of the grid-search machinery, cross-validation procedure, and downstream inspection identical; after tuning, predictions and probability outputs would be obtained from tuned_model_2_b in the same way the notebook produced y_pred_1_b and y_proba_1_b for the first tuned estimator.
Confusion Matrix
In [25] Copy
# Make predictions
y_pred_2_b=tuned_model_2_b.predict(X_test_2)
# create confustion matrix
fig, ax=plt.subplots()
sns.heatmap(pd.DataFrame(metrics.confusion_matrix(y_test_2, y_pred_2_b)), annot=True, cmap="Blues" ,fmt='g')
plt.title('Confusion matrix'); plt.ylabel('Actual label'); plt.xlabel('Predicted label')
ax.xaxis.set_ticklabels(['Down', 'Up']); ax.yaxis.set_ticklabels(['Down', 'Up'])
print("Accuracy:",metrics.accuracy_score(y_test_2, y_pred_2_b))
print("Precision:",metrics.precision_score(y_test_2, y_pred_2_b))
print("Recall:",metrics.recall_score(y_test_2, y_pred_2_b))Output
[stdout]
Accuracy: 0.6606765327695561
Precision: 0.6607142857142857
Recall: 0.9720853858784894In the notebookâs evaluation stage, the tuned_model_2_b estimator is applied to the temporally heldâout, windowed test set X_test_2 to produce y_pred_2_b, a sequence of discrete class labels the model assigns to each test example; those test examples were created earlier when framing historical price series into short sequences for the LSTM and classical models. The resulting label vector y_pred_2_b is then compared against the true labels y_test_2 via metrics.confusion_matrix to produce the confusion matrix that gets visualized with a seaborn heatmap (axis tick labels set to Down and Up), and the same predicted labels are used to compute and print standard classification summaries â accuracy, precision, and recall. This follows the same evaluation pattern used for tuned_model_1_b, tuned_model_3_b, and tuned_model_4_b (only the model and test partition differ), and complements earlier use of predict_proba for tuned_model_1_b where probability estimates were obtained instead of hard class labels.
ROC Curve
In [26] Copy
y_proba_2_b=tuned_model_2_b.predict_proba(X_test_2)[:, 1]
fpr, tpr, _=metrics.roc_curve(y_test_2, y_proba_2_b)
auc=metrics.roc_auc_score(y_test_2, y_proba_2_b)
plt.plot(fpr,tpr,label="AUC="+str(auc))
plt.legend(loc=4)
plt.plot([0, 1], [0, 1], linestyle='--') # plot no skill
plt.title('ROC-Curve')
plt.show()Output
The tuned_model_2_b estimator is asked for class membership probabilities on the temporally heldâout, windowed test set X_test_2, producing y_proba_2_b as the continuous score for the positive class (class index 1). Those probability scores are used instead of the earlier discrete labels stored in y_pred_2_b so the notebook can compute a full ROC curve and an AUC value that measure ranking performance across thresholds rather than a single hard decision. tuned_model_2_b is the pipeline_b estimator that resulted from the hyperparameter search where pipeline_b explored the small iterations_2_b budget, so the probabilities reflect the tuned classifier configuration found during model selection. In the notebookâs evaluation cell that began at gap_L638_638, y_proba_2_b flows into metrics.roc_curve and metrics.roc_auc_score to produce the false positive / true positive rates and the AUC used for plotting, and this pattern is repeated for other experiments (tuned_model_1_b on X_test_1, tuned_model_2_lstm on X_test_2_lstm, tuned_model_7_b on X_test_7) where the same probabilityâROCâAUC workflow is applied to different test splits and estimators.
LSTM
In [27] Copy
start=time.time()
# number of epochs
epochs=1
# number of units
LSTM_units_2_lstm=180
# number of samples
num_samples=1
# time_steps
look_back=1
# numer of features
num_features_2_lstm=X_train_2.shape[1]
# Regularization
dropout_rate=0.
recurrent_dropout=0.4
# print
verbose=0
#hyperparameter
batch_size=[1]
# hyperparameter
hyperparameter_2_lstm={'batch_size':batch_size}
# create Classifier
clf_2_lstm=KerasClassifier(build_fn=create_shallow_LSTM,
epochs=epochs,
LSTM_units=LSTM_units_2_lstm,
num_samples=num_samples,
look_back=look_back,
num_features=num_features_2_lstm,
dropout_rate=dropout_rate,
recurrent_dropout=recurrent_dropout,
verbose=verbose)
# Gridsearch
search_2_lstm=GridSearchCV(estimator=clf_2_lstm,
param_grid=hyperparameter_2_lstm,
n_jobs=-1,
cv=tscv,
scoring=scoring_lstm, # accuracy
refit=True,
return_train_score=False)
# Fit model
tuned_model_2_lstm=search_2_lstm.fit(X_train_2_lstm, y_train_2, shuffle=False, callbacks=[reset])
print("\n")
# View Accuracy
print(scoring_lstm +' of the best model: ', tuned_model_2_lstm.best_score_)
# best_score_ Mean cross-validated score of the best_estimator
print("\n")
# View best hyperparameters
print("Best hyperparameters:")
print('epochs:', tuned_model_2_lstm.best_estimator_.get_params()['epochs'])
print('batch_size:', tuned_model_2_lstm.best_estimator_.get_params()['batch_size'])
print('dropout_rate:', tuned_model_2_lstm.best_estimator_.get_params()['dropout_rate'])
print('recurrent_dropout:', tuned_model_2_lstm.best_estimator_.get_params()['recurrent_dropout'])
end=time.time()
print("\n")
print("Running Time:", end - start)Output
[stdout]
accuracy of the best model: 0.5562169312169312
Best hyperparameters:
epochs: 1
batch_size: 1
dropout_rate: 0.0
recurrent_dropout: 0.4
Running Time: 203.3864288330078The statement assigns the current wallâclock timestamp from Pythonâs time.time function to the variable start immediately before the LSTM hyperparameter setup and the GridSearchCV fit call, establishing a consistent timing anchor for the experiment that follows. In the context of the endâtoâend experimental notebook, that timestamp marks the beginning of the computational region whose duration the notebook records so the author can report the run time of the crossâvalidated LSTM training alongside the model quality metric (scoring_lstm and tuned_model_2_lstm.best_score_). The variable start is later paired with a corresponding end timestamp and a subtraction to produce the printed Running Time, which lets each LSTM fold (clf_1_lstm, clf_2_lstm, clf_3_lstm, clf_4_lstm, etc.) be compared not just by accuracy but by computational cost. The same start assignment pattern appears in the other LSTM experiment cells to provide a consistent timing protocol across different dataset splits and LSTM configurations, so the notebook can present both predictive performance and empirical training time as part of the model comparison.
Confusion Matrix
In [28] Copy
# Make predictions
y_pred_2_lstm=tuned_model_2_lstm.predict(X_test_2_lstm)
# create confustion matrix
fig, ax=plt.subplots()
sns.heatmap(pd.DataFrame(metrics.confusion_matrix(y_test_2, y_pred_2_lstm)), annot=True, cmap="Blues" ,fmt='g')
plt.title('Confusion matrix'); plt.ylabel('Actual label'); plt.xlabel('Predicted label')
ax.xaxis.set_ticklabels(['Down', 'Up']); ax.yaxis.set_ticklabels(['Down', 'Up'])
print("Accuracy:",metrics.accuracy_score(y_test_2, y_pred_2_lstm))
print("Precision:",metrics.precision_score(y_test_2, y_pred_2_lstm))
print("Recall:",metrics.recall_score(y_test_2, y_pred_2_lstm))Output
[stdout]
Accuracy: 0.6501057082452432
Precision: 0.6551339285714286
Recall: 0.9638752052545156The notebook applies the trained LSTM estimator for experiment 2 to the temporally held-out, windowed test set to produce predicted class labels for each sequence; those predictions are compared to the true labels for that same test partition to quantify how often the model forecasts an upward versus downward move. A confusion matrix is computed from the true-versus-predicted label pairs and rendered as a seaborn heatmap on a Matplotlib axes with the x/y ticks relabeled to the human-friendly classes Down and Up so you can visually inspect which class is being confused for the other. Finally, three scalar classification metricsâaccuracy, precision, and recallâare computed and printed from the same true and predicted arrays to give numeric summaries of overall correctness, positive prediction quality, and positive-class sensitivity. This evaluation step mirrors the pattern used for the other trained models (for example the experiment-1 and experiment-3 LSTMs and the classical pipeline whose y_pred_2_b was produced earlier) and serves the experiment notebookâs role of directly comparing LSTM performance against the classical learners on the same temporally consistent, windowed test data.
ROC Curve
In [29] Copy
y_proba_2_lstm=tuned_model_2_lstm.predict_proba(X_test_2_lstm)[:, 1]
fpr, tpr, _=metrics.roc_curve(y_test_2, y_proba_2_lstm)
auc=metrics.roc_auc_score(y_test_2, y_proba_2_lstm)
plt.plot(fpr,tpr,label="AUC="+str(auc))
plt.legend(loc=4)
plt.plot([0, 1], [0, 1], linestyle='--') # plot no skill
plt.title('ROC-Curve')
plt.show()Output
In the endâtoâend experiment notebook the pipeline named tuned_model_2_lstm is asked to emit class probabilities for the temporally heldâout, windowed test set X_test_2_lstm that was produced earlier when framing historical prices into sequence windows suitable for an LSTM; the predict_proba call returns a probability distribution over the two target classes and the notebook selects the probability assigned to the positive class as the continuous score for each test example. Those positiveâclass probabilities then feed into metrics.roc_curve together with the ground truth labels y_test_2 to produce false positive and true positive rates at many thresholds, and metrics.roc_auc_score summarizes the area under that ROC curve as a single scalar. The plotting calls render the ROC curve with the AUC shown in the legend, add a diagonal ânoâskillâ reference line, title the axes display as ROCâCurve, and render the Matplotlib figure to the output, producing the multiâaxes Figure object representation you see elsewhere in the notebook. This follows the same evaluation pattern used for tuned_model_1_lstm, tuned_model_2_b, and tuned_model_3_lstm, but differs in that tuned_model_2_lstm consumes the 3âD LSTM sequence input X_test_2_lstm while tuned_model_2_b operates on the flattened classical feature matrix X_test_2.
Model 3: Trading Volume
Baseline
In [30] Copy
# Model specific Parameter
# Number of iterations
#iterations_3_b=[20]
# Grid Search
# Regularization
#alpha_g_3_b=[0.00007, 0.00008, 0.00009]
#l1_ratio_g_3_b=[0., 0.2, 0.4, 0.6, 0.8, 1]
# Create hyperparameter options
#hyperparameters_g_3_b={'logistic__alpha':alpha_g_3_b,
# 'logistic__l1_ratio':l1_ratio_g_3_b,
# 'logistic__penalty':penalty_b,
# 'logistic__max_iter':iterations_3_b}
# Create grid search
#search_g_3_b=GridSearchCV(estimator=pipeline_b,
# param_grid=hyperparameters_g_3_b,
# cv=tscv,
# verbose=0,
# n_jobs=-1,
# scoring=scoring_b,
# refit=metric_b,
# return_train_score=False)
# Setting refit='Accuracy', refits an estimator on the whole dataset with the parameter setting that has the best cross-validated mean Accuracy score.
# For multiple metric evaluation, this needs to be a string denoting the scorer is used to find the best parameters for refitting the estimator at the end
# If return_train_score=True training results of CV will be saved as well
# Fit grid search
#tuned_model_3_b=search_g_3_b.fit(X_train_3, y_train_3)
#search_g_3_b.cv_results_
# Random Search
# Create regularization hyperparameter distribution using uniform distribution
alpha_r_3_b=uniform(loc=0.00001, scale=0.0001)
l1_ratio_r_3_b=uniform(loc=0, scale=1)
# Create hyperparameter options
hyperparameters_r_3_b={'logistic__alpha':alpha_r_3_b, 'logistic__l1_ratio':l1_ratio_r_3_b, 'logistic__penalty':penalty_b,'logistic__max_iter':iterations_3_b}
# Create randomized search
search_r_3_b=RandomizedSearchCV(pipeline_b, hyperparameters_r_3_b, n_iter=20, random_state=1, cv=tscv, verbose=0, n_jobs=-1, scoring=scoring_b, refit=metric_b, return_train_score=False)
# Setting refit='Accuracy', refits an estimator on the whole dataset with the parameter setting that has the best cross-validated Accuracy score.
# Fit randomized search
tuned_model_3_b=search_r_3_b.fit(X_train_3, y_train_3)
# View Cost function
print('Loss function:', tuned_model_3_b.best_estimator_.get_params()['logistic__loss'])
# View Accuracy
print(metric_b +' of the best model: ', tuned_model_3_b.best_score_);print("\n")
# best_score_ Mean cross-validated score of the best_estimator
# View best hyperparameters
print("Best hyperparameters:")
print('Number of iterations:', tuned_model_3_b.best_estimator_.get_params()['logistic__max_iter'])
print('Penalty:', tuned_model_3_b.best_estimator_.get_params()['logistic__penalty'])
print('Alpha:', tuned_model_3_b.best_estimator_.get_params()['logistic__alpha'])
print('l1_ratio:', tuned_model_3_b.best_estimator_.get_params()['logistic__l1_ratio'])
# Find the number of nonzero coefficients (selected features)
print("Total number of features:", len(tuned_model_3_b.best_estimator_.steps[1][1].coef_[0][:]))
print("Number of selected features:", np.count_nonzero(tuned_model_3_b.best_estimator_.steps[1][1].coef_[0][:]))
# Gridsearch table
plt.title('Gridsearch')
pvt_3_b=pd.pivot_table(pd.DataFrame(tuned_model_3_b.cv_results_), values='mean_test_accuracy', index='param_logistic__l1_ratio', columns='param_logistic__alpha')
ax_3_b=sns.heatmap(pvt_3_b, cmap="Blues")
plt.show()Output
[stdout]
Loss function: log
accuracy of the best model: 0.5521164021164021
Best hyperparameters:
Number of iterations: 20
Penalty: l1
Alpha: 1.836230045470262e-05
l1_ratio: 0.9168613345297285
Total number of features: 23
Number of selected features: 6Within the endâtoâend experimental notebook, alpha_r_3_b is created as a continuous uniform distribution object (from scipy.stats) that represents the sampling distribution for the logistic regression regularization strength used in the Trading Volume model workflow. Conceptually, this distribution defines the range of alpha values that RandomizedSearchCV will draw from when it explores hyperparameters for pipeline_b during the randomized search step that fits X_train_3 and y_train_3; later the chosen alpha value is reflected in tuned_model_3_b.best_estimator_.get_params()[’logistic__alpha’]. Numerically, the distribution is set to produce values starting at ten micro (1eâ5) up to about one hundred ten micro (â1.1eâ4), so the search focuses on very small, fineâgrained regularization strengths. The pattern mirrors other model blocks in the notebook that also create alpha_*_b as either explicit lists for GridSearchCV or as uniform distributions for RandomizedSearchCV, but the specific loc and scale here give a much narrower and lowerâmagnitude range than the distributions used in models like Model 7 or Model 6, which used larger loc/scale settings; the surrounding code pairs this alpha distribution with an l1_ratio distribution and passes both in the hyperparameter dictionary to RandomizedSearchCV so the elasticânet mixing and penalty strength are sampled jointly during crossâvalidated tuning.
Confusion Matrix
In [31] Copy
# Make predictions
y_pred_3_b=tuned_model_3_b.predict(X_test_3)
# create confustion matrix
fig, ax=plt.subplots()
sns.heatmap(pd.DataFrame(metrics.confusion_matrix(y_test_3, y_pred_3_b)), annot=True, cmap="Blues" ,fmt='g')
plt.title('Confusion matrix'); plt.ylabel('Actual label'); plt.xlabel('Predicted label')
ax.xaxis.set_ticklabels(['Down', 'Up']); ax.yaxis.set_ticklabels(['Down', 'Up'])
print("Accuracy:",metrics.accuracy_score(y_test_3, y_pred_3_b))
print("Precision:",metrics.precision_score(y_test_3, y_pred_3_b))
print("Recall:",metrics.recall_score(y_test_3, y_pred_3_b))Output
[stdout]
Accuracy: 0.6395348837209303
Precision: 0.6434689507494646
Recall: 0.986863711001642The statement calls the predict method on tuned_model_3_b to produce y_pred_3_b, so the tuned_model_3_b instance â the tuned classifier that was trained earlier in the notebook for experiment “3_b” â consumes the temporally heldâout, windowed test inputs contained in X_test_3 and emits discrete class labels for each test example. Those predicted labels then flow into the downstream evaluation steps: they are compared against the ground truth y_test_3 via metrics.confusion_matrix to populate the plotted confusion matrix and are used to compute scalar performance summaries (accuracy, precision, recall). This mirrors the same pattern repeated for the other classical-model experiments (y_pred_1_b, y_pred_2_b, y_pred_4_b) and contrasts with the LSTM example you saw earlier where tuned_model_2_lstm was asked for class probabilities via predict_proba instead of hard labels.
ROC Curve
In [32] Copy
y_proba_3_b=tuned_model_3_b.predict_proba(X_test_3)[:, 1]
fpr, tpr, _=metrics.roc_curve(y_test_3, y_proba_3_b)
auc=metrics.roc_auc_score(y_test_3, y_proba_3_b)
plt.plot(fpr,tpr,label="AUC="+str(auc))
plt.legend(loc=4)
plt.plot([0, 1], [0, 1], linestyle='--') # plot no skill
plt.title('ROC-Curve')
plt.show()Output
The line calls predict_proba on tuned_model_3_b to ask the fitted, tuned classifier for a probability distribution over the two target classes for each example in the temporally heldâout test set X_test_3 produced earlier in the pipeline; it then takes the probability assigned to the positive class as a continuous score for each test window so the modelâs probabilistic confidence can be used for ranking. Remember the predict_proba usage you saw with tuned_model_2_lstm â this is the same pattern applied to the third baseline model rather than the LSTM. Those positiveâclass scores are fed into metrics.roc_curve to produce false positive and true positive rate arrays across score thresholds and into metrics.roc_auc_score to produce the single AUC summary; the resulting FPR/TPR arrays are plotted as the ROC curve, the plot is annotated with the computed AUC in the legend placed at the lower right, a dashed diagonal is drawn to indicate noâskill performance, and the figure is titled and shown so you can visually compare discrimination performance for this model against the other workflows.
LSTM
In [33] Copy
start=time.time()
# number of epochs
epochs=1
# number of units
LSTM_units_3_lstm=180
# number of samples
num_samples=1
# time_steps
look_back=1
# numer of features
num_features_3_lstm=X_train_3.shape[1]
# Regularization
dropout_rate=0.
recurrent_dropout=0.4
# print
verbose=0
#hyperparameter
batch_size=[1]
# hyperparameter
hyperparameter_3_lstm={'batch_size':batch_size}
# create Classifier
clf_3_lstm=KerasClassifier(build_fn=create_shallow_LSTM,
epochs=epochs,
LSTM_units=LSTM_units_3_lstm,
num_samples=num_samples,
look_back=look_back,
num_features=num_features_3_lstm,
dropout_rate=dropout_rate,
recurrent_dropout=recurrent_dropout,
verbose=verbose)
# Gridsearch
search_3_lstm=GridSearchCV(estimator=clf_3_lstm,
param_grid=hyperparameter_3_lstm,
n_jobs=-1,
cv=tscv,
scoring=scoring_lstm, # accuracy
refit=True,
return_train_score=False)
# Fit model
tuned_model_3_lstm=search_3_lstm.fit(X_train_3_lstm, y_train_3, shuffle=False, callbacks=[reset])
print("\n")
# View Accuracy
print(scoring_lstm +' of the best model: ', tuned_model_3_lstm.best_score_)
# best_score_ Mean cross-validated score of the best_estimator
print("\n")
# View best hyperparameters
print("Best hyperparameters:")
print('epochs:', tuned_model_3_lstm.best_estimator_.get_params()['epochs'])
print('batch_size:', tuned_model_3_lstm.best_estimator_.get_params()['batch_size'])
print('dropout_rate:', tuned_model_3_lstm.best_estimator_.get_params()['dropout_rate'])
print('recurrent_dropout:', tuned_model_3_lstm.best_estimator_.get_params()['recurrent_dropout'])
end=time.time()
print("\n")
print("Running Time:", end - start)Output
[stdout]
accuracy of the best model: 0.56494708994709
Best hyperparameters:
epochs: 1
batch_size: 1
dropout_rate: 0.0
recurrent_dropout: 0.4
Running Time: 188.86053204536438The assignment of start to the current timestamp obtained from the time module marks the wallâclock beginning of the LSTM training experiment so the notebook can report how long the heavy operations take. In this cell start is set immediately before constructing the KerasClassifier, running GridSearchCV and calling the fit method, and a matching end timestamp is taken after fit completes; the difference between end and start is printed as Running Time, so the recorded value represents the elapsed real time for model construction, crossâvalidation folds and training (including any parallel work governed by the GridSearchCV n_jobs setting and the callbacks used during fit). This timing pattern is repeated across the other LSTM experiment cells so each tuned_model_* run produces a comparable perâmodel runtime measurement that the endâtoâend pipeline can use to compare computational cost alongside predictive performance.
Confusion Matrix
In [34] Copy
# Make predictions
y_pred_3_lstm=tuned_model_3_lstm.predict(X_test_3_lstm)
# create confustion matrix
fig, ax=plt.subplots()
sns.heatmap(pd.DataFrame(metrics.confusion_matrix(y_test_3, y_pred_3_lstm)), annot=True, cmap="Blues" ,fmt='g')
plt.title('Confusion matrix'); plt.ylabel('Actual label'); plt.xlabel('Predicted label')
ax.xaxis.set_ticklabels(['Down', 'Up']); ax.yaxis.set_ticklabels(['Down', 'Up'])
print("Accuracy:",metrics.accuracy_score(y_test_3, y_pred_3_lstm))
print("Precision:",metrics.precision_score(y_test_3, y_pred_3_lstm))
print("Recall:",metrics.recall_score(y_test_3, y_pred_3_lstm))Output
[stdout]
Accuracy: 0.6511627906976745
Precision: 0.6511375947995667
Recall: 0.986863711001642The notebook asks tuned_model_3_lstm to produce discrete class predictions on the temporally heldâout, windowed test set X_test_3_lstm â the same kind of sequence windows that were created earlier when framing historical prices for LSTM input â and then evaluates those predictions. It passes the predicted labels and the ground truth y_test_3 into sklearn.metrics.confusion_matrix to build the contingency table, renders that table as a seaborn heatmap on an axes returned by plt.subplots so the analyst can visually inspect true vs predicted counts, and relabels the plot ticks to the domain semantics ‘Down’ and ‘Up’. After the visual check, the cell computes and prints three common classification summaries (accuracy, precision, recall) using the corresponding metrics functions so you get scalar performance numbers that quantify overall correctness and the balance between false positives and false negatives. This follows the same evaluation pattern used elsewhere in the notebook (for example the tuned_model_3_b and the other tuned_model_* LSTM variants), with the key operational difference being that earlier, for tuned_model_2_lstm, predict_proba was used to obtain continuous positiveâclass scores for probabilistic/ROC-style analysis while here predict is used to obtain hard class labels for confusionâmatrix and thresholded metrics.
ROC Curve
In [35] Copy
y_proba_3_lstm=tuned_model_3_lstm.predict_proba(X_test_3_lstm)[:, 1]
fpr, tpr, _=metrics.roc_curve(y_test_3, y_proba_3_lstm)
auc=metrics.roc_auc_score(y_test_3, y_proba_3_lstm)
plt.plot(fpr,tpr,label="AUC="+str(auc))
plt.legend(loc=4)
plt.plot([0, 1], [0, 1], linestyle='--') # plot no skill
plt.title('ROC-Curve')
plt.show()Output
As with y_proba_2_lstm earlier, the notebook asks tuned_model_3_lstm to emit class probabilities for the temporally heldâout, windowed test set X_test_3_lstm; the evaluation selects the probability assigned to the positive class as the continuous score for each test example and pairs those scores with the true labels y_test_3. The sklearn.metrics.roc_curve call converts those paired scores and labels into a sequence of false positive and true positive rates across decision thresholds, and sklearn.metrics.roc_auc_score reduces the same inputs to a single areaâunderâtheâROC value that quantifies overall discrimination. Matplotlib is then used to draw the ROC trace, annotate it with the AUC in the legend, add the diagonal âno skillâ reference line, and render the figure to the output. This follows the same evaluation-and-plotting pattern used elsewhere in the notebook (for example with tuned_model_3_b, tuned_model_1_lstm, and tuned_model_4_lstm), differing only by which trained model and which temporally heldâout test split are being evaluated.
Model 4: Volatility + Return
In [36] Copy
# Model specific Parameter
# Number of iterations
iterations_4_b=[8]
# Grid Search
# Regularization
alpha_g_4_b=[0.0011, 0.0012, 0.0013]
l1_ratio_g_4_b=[0, 0.2, 0.4, 0.6, 0.8, 1]
# Create hyperparameter options
hyperparameters_g_4_b={'logistic__alpha':alpha_g_4_b,
'logistic__l1_ratio':l1_ratio_g_4_b,
'logistic__penalty':penalty_b,
'logistic__max_iter':iterations_4_b}
# Create grid search
search_g_4_b=GridSearchCV(estimator=pipeline_b,
param_grid=hyperparameters_g_4_b,
cv=tscv,
verbose=0,
n_jobs=-1,
scoring=scoring_b,
refit=metric_b,
return_train_score=False)
# Setting refit='Accuracy', refits an estimator on the whole dataset with the parameter setting that has the best cross-validated mean Accuracy score.
# For multiple metric evaluation, this needs to be a string denoting the scorer is used to find the best parameters for refitting the estimator at the end
# If return_train_score=True training results of CV will be saved as well
# Fit grid search
tuned_model_4_b=search_g_4_b.fit(X_train_4, y_train_4)
#search_g_4_b.cv_results_
# Random Search
# Create regularization hyperparameter distribution using uniform distribution
#alpha_r_4_b=uniform(loc=0.00006, scale=0.002) #loc=0.00006, scale=0.002
#l1_ratio_r_4_b=uniform(loc=0, scale=1)
# Create hyperparameter options
#hyperparameters_r_4_b={'logistic__alpha':alpha_r_4_b, 'logistic__l1_ratio':l1_ratio_r_4_b, 'logistic__penalty':penalty_b,'logistic__max_iter':iterations_4_b}
# Create randomized search
#search_r_4_b=RandomizedSearchCV(pipeline_b, hyperparameters_r_4_b, n_iter=10, random_state=1, cv=tscv, verbose=0, n_jobs=-1, scoring=scoring_b, refit=metric_b, return_train_score=False)
# Setting refit='Accuracy', refits an estimator on the whole dataset with the parameter setting that has the best cross-validated Accuracy score.
# Fit randomized search
#tuned_model_4_b=search_r_4_b.fit(X_train_4, y_train_4)
# View Cost function
print('Loss function:', tuned_model_4_b.best_estimator_.get_params()['logistic__loss'])
# View Accuracy
print(metric_b +' of the best model: ', tuned_model_4_b.best_score_);print("\n")
# best_score_ Mean cross-validated score of the best_estimator
# View best hyperparameters
print("Best hyperparameters:")
print('Number of iterations:', tuned_model_4_b.best_estimator_.get_params()['logistic__max_iter'])
print('Penalty:', tuned_model_4_b.best_estimator_.get_params()['logistic__penalty'])
print('Alpha:', tuned_model_4_b.best_estimator_.get_params()['logistic__alpha'])
print('l1_ratio:', tuned_model_4_b.best_estimator_.get_params()['logistic__l1_ratio'])
# Find the number of nonzero coefficients (selected features)
print("Total number of features:", len(tuned_model_4_b.best_estimator_.steps[1][1].coef_[0][:]))
print("Number of selected features:", np.count_nonzero(tuned_model_4_b.best_estimator_.steps[1][1].coef_[0][:]))
# Gridsearch table
plt.title('Gridsearch')
pvt_4_b=pd.pivot_table(pd.DataFrame(tuned_model_4_b.cv_results_), values='mean_test_accuracy', index='param_logistic__l1_ratio', columns='param_logistic__alpha')
ax_4_b=sns.heatmap(pvt_4_b, cmap="Blues")
plt.show()Output
[stdout]
Loss function: log
accuracy of the best model: 0.5206349206349207
Best hyperparameters:
Number of iterations: 8
Penalty: elasticnet
Alpha: 0.0012
l1_ratio: 0.6
Total number of features: 46
Number of selected features: 19iterations_4_b is a single-element list containing the integer eight; in the grid search that follows it provides the candidate value for the logistic regression solver’s maximum-iteration hyperparameter by being included in hyperparameters_g_4_b under the logistic__max_iter key. Placing the iteration budget in a list is necessary because GridSearchCV expects iterables for each hyperparameter, so iterations_4_b makes max_iter an explicit (but non-varying) dimension of the param grid used when pipeline_b is cross-validated with tscv and scoring_b and ultimately refit according to metric_b, producing tuned_model_4_b after calling fit on X_train_4 and y_train_4. Because only one value is supplied, the grid search will not sweep over different max_iter values but preserves the same parameterization pattern used across the notebookâs baseline logistic pipelines, and later code inspects the chosen max_iter via the best_estimator_ returned by GridSearchCV. The randomized-search alternative in this cell would also reference iterations_4_b if enabled, and the pattern mirrors other model cells where iterations_1_b, iterations_5_b and iterations_7_b play the same role but with different numeric choices (for example some models use ten instead of eight), reflecting per-model iteration budgets used during the classical learner experiments that are being compared against the LSTM pipeline.
Confusion Matrix
In [37] Copy
# Make predictions
y_pred_4_b=tuned_model_4_b.predict(X_test_4)
# create confustion matrix
fig, ax=plt.subplots()
sns.heatmap(pd.DataFrame(metrics.confusion_matrix(y_test_4, y_pred_4_b)), annot=True, cmap="Blues" ,fmt='g')
plt.title('Confusion matrix'); plt.ylabel('Actual label'); plt.xlabel('Predicted label')
ax.xaxis.set_ticklabels(['Down', 'Up']); ax.yaxis.set_ticklabels(['Down', 'Up'])
print("Accuracy:",metrics.accuracy_score(y_test_4, y_pred_4_b))
print("Precision:",metrics.precision_score(y_test_4, y_pred_4_b))
print("Recall:",metrics.recall_score(y_test_4, y_pred_4_b))Output
[stdout]
Accuracy: 0.6004228329809725
Precision: 0.6501950585175552
Recall: 0.8210180623973727The notebook asks the tuned_model_4_b instance to generate class predictions for the temporally heldâout test set X_test_4 so we can evaluate how that particular tuned classical pipeline performs on unseen data; X_test_4 contains the preprocessed, windowed or featureâengineered examples produced earlier in the pipeline and the resulting discrete outputs are the modelâs estimated up/down labels for each test time window. Those predicted labels are then paired with the true labels y_test_4 to produce the confusion matrix and derive accuracy, precision and recall for this experiment. This call follows the same evaluation pattern used for tuned_model_1_b, tuned_model_2_b and tuned_model_3_b, and mirrors how the LSTM pipeline produced discrete predictions for its windowed test set, enabling a direct, temporally consistent comparison between the classical learner represented by tuned_model_4_b and the other models in the endâtoâend forecasting experiment.
ROC Curve
In [38] Copy
y_proba_4_b=tuned_model_4_b.predict_proba(X_test_4)[:, 1]
fpr, tpr, _=metrics.roc_curve(y_test_4, y_proba_4_b)
auc=metrics.roc_auc_score(y_test_4, y_proba_4_b)
plt.plot(fpr,tpr,label="AUC="+str(auc))
plt.legend(loc=4)
plt.plot([0, 1], [0, 1], linestyle='--') # plot no skill
plt.title('ROC-Curve')
plt.show()Output
Like y_proba_3_lstm earlier, the notebook asks tuned_model_4_b to produce class probabilities for the temporally heldâout, windowed test set X_test_4; it selects the probability assigned to the positive class for each test example and passes those continuous scores into scikitâlearn’s ROC utilities (metrics.roc_curve and metrics.roc_auc_score) to compute false positive and true positive rates and the area under the ROC curve. The notebook then plots the computed fpr/tpr curve, annotates the plot with the AUC in the legend, overlays a diagonal noâskill reference line, and displays the figure so you can visually and numerically assess how well the Model 4 baseline discriminates the target on the heldâout windows, following the same evaluation pattern used for the other model variants in the experiment.
LSTM
In [39] Copy
start=time.time()
# number of epochs
epochs=1
# number of units
LSTM_units_4_lstm=200
# number of samples
num_samples=1
# time_steps
look_back=1
# numer of features
num_features_4_lstm=X_train_4.shape[1]
# Regularization
dropout_rate=0.
recurrent_dropout=0.4
# print
verbose=0
#hyperparameter
batch_size=[1]
# hyperparameter
hyperparameter_4_lstm={'batch_size':batch_size}
# create Classifier
clf_4_lstm=KerasClassifier(build_fn=create_shallow_LSTM,
epochs=epochs,
LSTM_units=LSTM_units_4_lstm,
num_samples=num_samples,
look_back=look_back,
num_features=num_features_4_lstm,
dropout_rate=dropout_rate,
recurrent_dropout=recurrent_dropout,
verbose=verbose)
# Gridsearch
search_4_lstm=GridSearchCV(estimator=clf_4_lstm,
param_grid=hyperparameter_4_lstm,
n_jobs=-1,
cv=tscv,
scoring=scoring_lstm, # accuracy
refit=True,
return_train_score=False)
# Fit model
tuned_model_4_lstm=search_4_lstm.fit(X_train_4_lstm, y_train_4, shuffle=False, callbacks=[reset])
print("\n")
# View Accuracy
print(scoring_lstm +' of the best model: ', tuned_model_4_lstm.best_score_)
# best_score_ Mean cross-validated score of the best_estimator
print("\n")
# View best hyperparameters
print("Best hyperparameters:")
print('epochs:', tuned_model_4_lstm.best_estimator_.get_params()['epochs'])
print('batch_size:', tuned_model_4_lstm.best_estimator_.get_params()['batch_size'])
print('dropout_rate:', tuned_model_4_lstm.best_estimator_.get_params()['dropout_rate'])
print('recurrent_dropout:', tuned_model_4_lstm.best_estimator_.get_params()['recurrent_dropout'])
end=time.time()
print("\n")
print("Running Time:", end - start)Output
[stdout]
accuracy of the best model: 0.5441798941798942
Best hyperparameters:
epochs: 1
batch_size: 1
dropout_rate: 0.0
recurrent_dropout: 0.4
Running Time: 244.36903405189514At the start of the LSTM training cell the notebook captures a wallâclock timestamp into the variable named start by calling the time.time function. That timestamp is used to bracket the expensive work that follows â building the KerasClassifier with create_shallow_LSTM, running GridSearchCV with the timeâseries CV and scoring function, and fitting the tuned model â and later the notebook captures a second timestamp and prints their difference as the Running Time for this experiment. Recording elapsed time here provides an explicit, perâexperiment runtime measurement so the notebook can report and compare how long each LSTM training/gridâsearch run takes; the same timing pattern is applied in the other LSTM blocks to produce comparable timing statistics across model variants.
Confusion Matrix
In [40] Copy
# Make predictions
y_pred_4_lstm=tuned_model_4_lstm.predict(X_test_4_lstm)
# create confustion matrix
fig, ax=plt.subplots()
sns.heatmap(pd.DataFrame(metrics.confusion_matrix(y_test_4, y_pred_4_lstm)), annot=True, cmap="Blues" ,fmt='g')
plt.title('Confusion matrix'); plt.ylabel('Actual label'); plt.xlabel('Predicted label')
ax.xaxis.set_ticklabels(['Down', 'Up']); ax.yaxis.set_ticklabels(['Down', 'Up'])
print("Accuracy:",metrics.accuracy_score(y_test_4, y_pred_4_lstm))
print("Precision:",metrics.precision_score(y_test_4, y_pred_4_lstm))
print("Recall:",metrics.recall_score(y_test_4, y_pred_4_lstm))Output
[stdout]
Accuracy: 0.6331923890063424
Precision: 0.6714659685863874
Recall: 0.8423645320197044y_pred_4_lstm asks tuned_model_4_lstm to emit discrete class labels for the temporally heldâout, windowed LSTM test set X_test_4_lstm; those predictions are then compared against the true labels y_test_4 to produce a confusion matrix via metrics.confusion_matrix, which is visualized with a seaborn heatmap (wrapped as a DataFrame and annotated, using a blue color map) and annotated axes titled and ticked as Down/Up to reflect the binary movement classes. After rendering the matrix, the notebook computes and prints three scalar classification summariesâaccuracy, precision, and recallâusing metrics.accuracy_score, metrics.precision_score, and metrics.recall_score so the LSTM trained under the configuration labeled â4â can be directly compared to other learners. This follows the same evaluation pattern used for y_pred_1_lstm and y_pred_3_lstm (both LSTM variants on their respective windowed test sets) and mirrors the presentation used for tuned_model_4_b (a nonâsequence/classical model evaluated on X_test_4), ensuring a consistent visual and numeric comparison across model architectures and feature/target pipelines in the experiment.
ROC Curve
In [41] Copy
y_proba_4_lstm=tuned_model_4_lstm.predict_proba(X_test_4_lstm)[:, 1]
fpr, tpr, _=metrics.roc_curve(y_test_4, y_proba_4_lstm)
auc=metrics.roc_auc_score(y_test_4, y_proba_4_lstm)
plt.plot(fpr,tpr,label="AUC="+str(auc))
plt.legend(loc=4)
plt.plot([0, 1], [0, 1], linestyle='--') # plot no skill
plt.title('ROC-Curve')
plt.show()Output
Within the experiment notebook, the call that asks tuned_model_4_lstm for class probabilities on X_test_4_lstm converts the trained LSTM’s outputs for each windowed test sample into a numeric score representing the model’s estimated probability of the positive movement class (the “Up” label) so those soft outputs can be evaluated across decision thresholds rather than as a single hard label; remember the LSTM expects its input shaped as samples, time steps, and features so X_test_4_lstm is the temporally framed 3âD test set. Those probability scores are paired with the true binary labels y_test_4 and fed into metrics.roc_curve to produce the false positive and true positive rates across thresholds, and into metrics.roc_auc_score to compute the single summary AUC value that quantifies discriminative ability. The subsequent plotting routine draws the ROC curve, annotates it with the AUC in the legend (placed in the plot’s lowerâright), and overlays the diagonal noâskill reference line so you can visually compare the model’s tradeoff between sensitivity and false alarms. This pattern is the same evaluation idiom used elsewhere in the notebook for other trained estimators (for example the tuned_model_1_lstm, tuned_model_3_lstm and the classical tuned_model_4_b), differing only in which fitted model and which temporally heldâout test split are being scored; earlier you used discrete predictions from tuned_model_4_lstm to build a confusion matrix, whereas here you use probabilistic outputs to generate the ROC curve and AUC.
Model 5: Volatility + Return
Baseline
In [42] Copy
# Model specific Parameter
# Number of iterations
iterations_5_b=[10]
# Grid Search
# Regularization
alpha_g_5_b=[0.0001, 0.0003, 0.0005]
l1_ratio_g_5_b=[0, 0.2, 0.4, 0.6, 0.8, 1]
# Create hyperparameter options
hyperparameters_g_5_b={'logistic__alpha':alpha_g_5_b,
'logistic__l1_ratio':l1_ratio_g_5_b,
'logistic__penalty':penalty_b,
'logistic__max_iter':iterations_5_b}
# Create grid search
search_g_5_b=GridSearchCV(estimator=pipeline_b,
param_grid=hyperparameters_g_5_b,
cv=tscv,
verbose=0,
n_jobs=-1,
scoring=scoring_b,
refit=metric_b,
return_train_score=False)
# Setting refit='Accuracy', refits an estimator on the whole dataset with the parameter setting that has the best cross-validated mean Accuracy score.
# For multiple metric evaluation, this needs to be a string denoting the scorer is used to find the best parameters for refitting the estimator at the end
# If return_train_score=True training results of CV will be saved as well
# Fit grid search
tuned_model_5_b=search_g_5_b.fit(X_train_5, y_train_5)
#search_g_5_b.cv_results_
# Random Search
# Create regularization hyperparameter distribution using uniform distribution
#alpha_r_5_b=uniform(loc=0.00006, scale=0.002) #loc=0.00006, scale=0.002
#l1_ratio_r_5_b=uniform(loc=0, scale=1)
# Create hyperparameter options
#hyperparameters_r_5_b={'logistic__alpha':alpha_r_5_b, 'logistic__l1_ratio':l1_ratio_r_5_b, 'logistic__penalty':penalty_b,'logistic__max_iter':iterations_5_b}
# Create randomized search
#search_r_5_b=RandomizedSearchCV(pipeline_b, hyperparameters_r_5_b, n_iter=10, random_state=1, cv=tscv, verbose=0, n_jobs=-1, scoring=scoring_b, refit=metric_b, return_train_score=False)
# Setting refit='Accuracy', refits an estimator on the whole dataset with the parameter setting that has the best cross-validated Accuracy score.
# Fit randomized search
#tuned_model_5_b=search_r_4_b.fit(X_train_5, y_train_5)
# View Cost function
print('Loss function:', tuned_model_5_b.best_estimator_.get_params()['logistic__loss'])
# View Accuracy
print(metric_b +' of the best model: ', tuned_model_5_b.best_score_);print("\n")
# best_score_ Mean cross-validated score of the best_estimator
# View best hyperparameters
print("Best hyperparameters:")
print('Number of iterations:', tuned_model_5_b.best_estimator_.get_params()['logistic__max_iter'])
print('Penalty:', tuned_model_5_b.best_estimator_.get_params()['logistic__penalty'])
print('Alpha:', tuned_model_5_b.best_estimator_.get_params()['logistic__alpha'])
print('l1_ratio:', tuned_model_5_b.best_estimator_.get_params()['logistic__l1_ratio'])
# Find the number of nonzero coefficients (selected features)
print("Total number of features:", len(tuned_model_5_b.best_estimator_.steps[1][1].coef_[0][:]))
print("Number of selected features:", np.count_nonzero(tuned_model_5_b.best_estimator_.steps[1][1].coef_[0][:]))
# Gridsearch table
plt.title('Gridsearch')
pvt_5_b=pd.pivot_table(pd.DataFrame(tuned_model_5_b.cv_results_), values='mean_test_accuracy', index='param_logistic__l1_ratio', columns='param_logistic__alpha')
ax_5_b=sns.heatmap(pvt_5_b, cmap="Blues")
plt.show()Output
[stdout]
Loss function: log
accuracy of the best model: 0.5551587301587302
Best hyperparameters:
Number of iterations: 10
Penalty: elasticnet
Alpha: 0.0005
l1_ratio: 0.6
Total number of features: 46
Number of selected features: 15iterations_5_b is the singleâelement list that supplies candidate values for the logistic estimator’s maximum iteration budget when tuning Model 5 (the Volatility + Return baseline) â it gets bundled into the hyperparameter dictionary used by GridSearchCV so that the pipeline step named pipeline_b will be trained with that maxâiteration setting during each crossâvalidation trial. In the notebook’s tuning flow the value in iterations_5_b flows into hyperparameters_g_5_b, which GridSearchCV consumes (alongside alpha and l1_ratio grids and the penalty choices) together with the timeâseries crossâvalidator tscv, the scoring function scoring_b and the refit metric metric_b; calling search_g_5_b.fit on X_train_5 and y_train_5 produces tuned_model_5_b that is later inspected for loss, best_score_ and best hyperparameters. The same pattern appears for other model blocks (for example iterations_4_b and iterations_1_b use a different numeric candidate), so iterations_5_b follows the projectâs repeated convention of exposing the logistic__max_iter parameter as a oneâitem list for inclusion in the grid; the randomized search variants shown in comments would also reference this iterations list if they were enabled.
Confusion Matrix
In [43] Copy
# Make predictions
y_pred_5_b=tuned_model_5_b.predict(X_test_5)
# create confustion matrix
fig, ax=plt.subplots()
sns.heatmap(pd.DataFrame(metrics.confusion_matrix(y_test_5, y_pred_5_b)), annot=True, cmap="Blues" ,fmt='g')
plt.title('Confusion matrix'); plt.ylabel('Actual label'); plt.xlabel('Predicted label')
ax.xaxis.set_ticklabels(['Down', 'Up']); ax.yaxis.set_ticklabels(['Down', 'Up'])
print("Accuracy:",metrics.accuracy_score(y_test_5, y_pred_5_b))
print("Precision:",metrics.precision_score(y_test_5, y_pred_5_b))
print("Recall:",metrics.recall_score(y_test_5, y_pred_5_b))Output
[stdout]
Accuracy: 0.5909090909090909
Precision: 0.629976580796253
Recall: 0.8834154351395731The line invokes the classifier instance named tuned_model_5_b to produce predicted binary movement labels for the temporally heldâout, windowed test set X_test_5; those predictions are the model’s discrete Up/Down calls on the engineered features that emerged from the pipeline’s preprocessing and windowing steps. Conceptually, the data flow here is feature matrix â tuned_model_5_b.predict â categorical predictions, which are then fed into a confusion matrix visualization and the scalar scorers that follow to quantify accuracy, precision and recall against the true labels y_test_5. This mirrors the pattern used elsewhere in the notebookâfor example the y_pred_4_b and y_pred_2_b baselines and the y_pred_5_lstm runâbut differs in that tuned_model_5_b is a tuned classical learner operating on a twoâdimensional samplesÃfeatures matrix, whereas the LSTM variant required the threeâdimensional samplesÃtimestepsÃfeatures shape noted earlier. The purpose in the endâtoâend experimental notebook is to collect these label outputs so the heatmap and printed metrics can directly compare forecasting performance between the classical tuned_model_5_b and the other tuned models.
ROC Curve
In [44] Copy
y_proba_5_b=tuned_model_5_b.predict_proba(X_test_5)[:, 1]
fpr, tpr, _=metrics.roc_curve(y_test_5, y_proba_5_b)
auc=metrics.roc_auc_score(y_test_5, y_proba_5_b)
plt.plot(fpr,tpr,label="AUC="+str(auc))
plt.legend(loc=4)
plt.plot([0, 1], [0, 1], linestyle='--') # plot no skill
plt.title('ROC-Curve')
plt.show()Output
At the evaluation stage the notebook asks tuned_model_5_b to produce probabilistic forecasts for every example in the temporally held-out test matrix X_test_5; tuned_model_5_b is the hyperparameterâtuned classical classifier trained earlier in the pipeline, and its predict_proba method emits a probability distribution over the two movement classes for each test window, from which the notebook selects the probability assigned to the positive (Up) class as a continuous score. Those perâsample Up probabilities are what the ROC machinery expects â they flow next into metrics.roc_curve and metrics.roc_auc_score to compute false positive and true positive rates and an AUC summary â a different output than the discrete labels produced by the LSTM call you saw earlier that assigned y_pred_4_lstm â and this call follows the same evaluation pattern used for the other classical and LSTM variants (for example the tuned_model_1_b, tuned_model_7_b and tuned_model_5_lstm cases), differing only by which tuned model and which heldâout test set are being scored.
LSTM
In [45] Copy
start=time.time()
# number of epochs
epochs=1
# number of units
LSTM_units_5_lstm=190
# number of samples
num_samples=1
# time_steps
look_back=1
# numer of features
num_features_5_lstm=X_train_5.shape[1]
# Regularization
dropout_rate=0.
recurrent_dropout=0.3
# print
verbose=0
#hyperparameter
batch_size=[1]
# hyperparameter
hyperparameter_5_lstm={'batch_size':batch_size}
# create Classifier
clf_5_lstm=KerasClassifier(build_fn=create_shallow_LSTM,
epochs=epochs,
LSTM_units=LSTM_units_5_lstm,
num_samples=num_samples,
look_back=look_back,
num_features=num_features_5_lstm,
dropout_rate=dropout_rate,
recurrent_dropout=recurrent_dropout,
verbose=verbose)
# Gridsearch
search_5_lstm=GridSearchCV(estimator=clf_5_lstm,
param_grid=hyperparameter_5_lstm,
n_jobs=-1,
cv=tscv,
scoring=scoring_lstm, # accuracy
refit=True,
return_train_score=False)
# Fit model
tuned_model_5_lstm=search_5_lstm.fit(X_train_5_lstm, y_train_5, shuffle=False, callbacks=[reset])
print("\n")
# View Accuracy
print(scoring_lstm +' of the best model: ', tuned_model_5_lstm.best_score_)
# best_score_ Mean cross-validated score of the best_estimator
print("\n")
# View best hyperparameters
print("Best hyperparameters:")
print('epochs:', tuned_model_5_lstm.best_estimator_.get_params()['epochs'])
print('batch_size:', tuned_model_5_lstm.best_estimator_.get_params()['batch_size'])
print('dropout_rate:', tuned_model_5_lstm.best_estimator_.get_params()['dropout_rate'])
print('recurrent_dropout:', tuned_model_5_lstm.best_estimator_.get_params()['recurrent_dropout'])
end=time.time()
print("\n")
print("Running Time:", end - start)Output
[stdout]
accuracy of the best model: 0.5933862433862434
Best hyperparameters:
epochs: 1
batch_size: 1
dropout_rate: 0.0
recurrent_dropout: 0.3
Running Time: 303.28468012809753The assignment of start to time.time acts as a simple wallâclock stopwatch kicked off immediately before assembling the KerasClassifier, executing the GridSearchCV and calling fit for this LSTM experiment so the notebook can later capture end and print the elapsed Running Time for the training and hyperparameter search. In the context of the endâtoâend experiment notebook that compares LSTM models and classical learners, that timing lets each LSTM cell record how long its training plus crossâvalidation took alongside the reported performance metrics (for example the best_score_ printed from tuned_model_5_lstm), which is important because the pipeline evaluates models both by forecasting accuracy and by computational cost. The same pattern appears in the other LSTM cells: start is placed just before model construction and fit, and an end timestamp is taken after fit to produce a perâmodel elapsed time, so runtime comparisons across the different lookbacks, unit counts and regularization settings are directly measurable.
Confusion Matrix
In [46] Copy
# Make predictions
y_pred_5_lstm=tuned_model_5_lstm.predict(X_test_5_lstm)
# create confustion matrix
fig, ax=plt.subplots()
sns.heatmap(pd.DataFrame(metrics.confusion_matrix(y_test_5, y_pred_5_lstm)), annot=True, cmap="Blues" ,fmt='g')
plt.title('Confusion matrix'); plt.ylabel('Actual label'); plt.xlabel('Predicted label')
ax.xaxis.set_ticklabels(['Down', 'Up']); ax.yaxis.set_ticklabels(['Down', 'Up'])
print("Accuracy:",metrics.accuracy_score(y_test_5, y_pred_5_lstm))
print("Precision:",metrics.precision_score(y_test_5, y_pred_5_lstm))
print("Recall:",metrics.recall_score(y_test_5, y_pred_5_lstm))Output
[stdout]
Accuracy: 0.6596194503171248
Precision: 0.662883087400681
Recall: 0.9589490968801314The assignment to y_pred_5_lstm asks the tuned_model_5_lstm to produce class predictions for every example in X_test_5_lstm, which is the temporally heldâout set of sequence windows formatted for the LSTM; those predicted labels are captured in y_pred_5_lstm and immediately compared to the true labels y_test_5 to quantify performance. The notebook then converts the raw counts from metrics.confusion_matrix into a tabular structure and renders it as a seaborn heatmap, annotating the cells and relabeling the axes to the two movement classes Down and Up so you can visually inspect false positives, false negatives and correct classifications. Finally the code computes and prints standard labelâbased metrics â accuracy, precision and recall â using the same metrics module to summarize the LSTMâs pointâprediction performance on the fiveâstep experiment. This follows the same evaluation pattern used elsewhere in the notebook for other models and horizons, but differs from the earlier y_proba_5_b flow in that tuned_model_5_lstm returns discrete class labels for sequenceâshaped inputs (X_test_5_lstm) rather than the probability vectors produced by the classical tuned_model_5_b on 2D feature matrices (X_test_5), so the resulting metrics are computed directly on predicted classes rather than on a thresholded probability score.
ROC Curce
In [47] Copy
y_proba_5_lstm=tuned_model_5_lstm.predict_proba(X_test_5_lstm)[:, 1]
fpr, tpr, _=metrics.roc_curve(y_test_5, y_proba_5_lstm)
auc=metrics.roc_auc_score(y_test_5, y_proba_5_lstm)
plt.plot(fpr,tpr,label="AUC="+str(auc))
plt.legend(loc=4)
plt.plot([0, 1], [0, 1], linestyle='--') # plot no skill
plt.title('ROC-Curve')
plt.show()Output
The cell requests probabilistic forecasts from tuned_model_5_lstm for the temporally held-out sequence test set X_test_5_lstm; it extracts the probability mass that the model assigns to the positive (Up) class and uses those continuous scores together with the true labels y_test_5 to compute the ROC operating points via metrics.roc_curve and a single-number summary via metrics.roc_auc_score. The false positive and true positive rate arrays are plotted as the ROC curve with the computed AUC shown in the legend, a diagonal no-skill reference line is overlaid, and the figure is titled and displayed. Functionally this is the LSTM-side equivalent of the earlier classical-model evaluation that produced y_proba_5_b from tuned_model_5_b, but it differs in that the input here is the sequence-framed test tensor appropriate for the LSTM architecture and the probabilities come from the hyperparameter-tuned LSTM we trained (the training run whose wall-clock was marked with start=time.time). The output provides a visual and numeric measure of how well the LSTMâs probability estimates discriminate Up versus Down on the held-out temporal test fold, supporting the notebookâs end-to-end comparison between recurrent and classical learners.
Model 6: Trading Volume + Return
Baseline
In [48] Copy
# Model specific Parameter
# Number of iterations
iterations_6_b=[8]
# Grid Search
# Regularization
alpha_g_6_b=[0.0011, 0.0012, 0.0013]
l1_ratio_g_6_b=[0, 0.2, 0.4, 0.6, 0.8, 1]
# Create hyperparameter options
hyperparameters_g_6_b={'logistic__alpha':alpha_g_6_b,
'logistic__l1_ratio':l1_ratio_g_6_b,
'logistic__penalty':penalty_b,
'logistic__max_iter':iterations_6_b}
# Create grid search
search_g_6_b=GridSearchCV(estimator=pipeline_b,
param_grid=hyperparameters_g_6_b,
cv=tscv,
verbose=0,
n_jobs=-1,
scoring=scoring_b,
refit=metric_b,
return_train_score=False)
# Setting refit='Accuracy', refits an estimator on the whole dataset with the parameter setting that has the best cross-validated mean Accuracy score.
# For multiple metric evaluation, this needs to be a string denoting the scorer is used to find the best parameters for refitting the estimator at the end
# If return_train_score=True training results of CV will be saved as well
# Fit grid search
tuned_model_6_b=search_g_6_b.fit(X_train_6, y_train_6)
#search_g_6_b.cv_results_
# Random Search
# Create regularization hyperparameter distribution using uniform distribution
#alpha_r_6_b=uniform(loc=0.00006, scale=0.002) #loc=0.00006, scale=0.002
#l1_ratio_r_6_b=uniform(loc=0, scale=1)
# Create hyperparameter options
#hyperparameters_r_6_b={'logistic__alpha':alpha_r_6_b, 'logistic__l1_ratio':l1_ratio_r_6_b, 'logistic__penalty':penalty_b,'logistic__max_iter':iterations_6_b}
# Create randomized search
#search_r_6_b=RandomizedSearchCV(pipeline_b, hyperparameters_r_6_b, n_iter=10, random_state=1, cv=tscv, verbose=0, n_jobs=-1, scoring=scoring_b, refit=metric_b, return_train_score=False)
# Setting refit='Accuracy', refits an estimator on the whole dataset with the parameter setting that has the best cross-validated Accuracy score.
# Fit randomized search
#tuned_model_6_b=search_r_4_b.fit(X_train_6, y_train_6)
# View Cost function
print('Loss function:', tuned_model_6_b.best_estimator_.get_params()['logistic__loss'])
# View Accuracy
print(metric_b +' of the best model: ', tuned_model_6_b.best_score_);print("\n")
# best_score_ Mean cross-validated score of the best_estimator
# View best hyperparameters
print("Best hyperparameters:")
print('Number of iterations:', tuned_model_6_b.best_estimator_.get_params()['logistic__max_iter'])
print('Penalty:', tuned_model_6_b.best_estimator_.get_params()['logistic__penalty'])
print('Alpha:', tuned_model_6_b.best_estimator_.get_params()['logistic__alpha'])
print('l1_ratio:', tuned_model_6_b.best_estimator_.get_params()['logistic__l1_ratio'])
# Find the number of nonzero coefficients (selected features)
print("Total number of features:", len(tuned_model_6_b.best_estimator_.steps[1][1].coef_[0][:]))
print("Number of selected features:", np.count_nonzero(tuned_model_6_b.best_estimator_.steps[1][1].coef_[0][:]))
# Gridsearch table
plt.title('Gridsearch')
pvt_6_b=pd.pivot_table(pd.DataFrame(tuned_model_6_b.cv_results_), values='mean_test_accuracy', index='param_logistic__l1_ratio', columns='param_logistic__alpha')
ax_6_b=sns.heatmap(pvt_6_b, cmap="Blues")
plt.show()Output
[stdout]
Loss function: log
accuracy of the best model: 0.5419312169312169
Best hyperparameters:
Number of iterations: 8
Penalty: l1
Alpha: 0.0013
l1_ratio: 0
Total number of features: 46
Number of selected features: 6iterations_6_b is the small, singleâelement list that supplies the candidate value for the logistic regression solver’s maximum iteration count when building the hyperparameter grid for the Model 6 experiment (the Trading Volume + Return baseline). Within the notebook’s endâtoâend pipeline, each model prepares a similar set of hyperparameter lists and then plugs them into a GridSearchCV or RandomizedSearchCV over the baseline estimator pipeline_b; iterations_6_b specifically becomes the value used for the logistic__max_iter entry in the hyperparameters_g_6_b mapping that GridSearchCV will iterate over. Because iterations_6_b contains only one value, GridSearchCV will hold the solver iteration budget fixed while it explores combinations of alpha, l1_ratio and penalty for X_train_6/y_train_6, and the chosen iterations value is carried through into the best_estimator_ that tuned_model_6_b yields after fitting. This follows the same pattern used by other model sections (for example iterations_4_b is the same single value and iterations_7_b is a different single value), so the notebook keeps the max_iter parameter explicit and modelâspecific even when it is not being swept across multiple options. The iterations setting therefore affects how many internal optimization steps the logistic step in pipeline_b is allowed during crossâvalidation and in the final refit that GridSearchCV performs, and its value is included in the run timing started earlier with start=time.time.
Confusion Matrix
In [49] Copy
# Make predictions
y_pred_6_b=tuned_model_6_b.predict(X_test_6)
# create confustion matrix
fig, ax=plt.subplots()
sns.heatmap(pd.DataFrame(metrics.confusion_matrix(y_test_6, y_pred_6_b)), annot=True, cmap="Blues" ,fmt='g')
plt.title('Confusion matrix'); plt.ylabel('Actual label'); plt.xlabel('Predicted label')
ax.xaxis.set_ticklabels(['Down', 'Up']); ax.yaxis.set_ticklabels(['Down', 'Up'])
print("Accuracy:",metrics.accuracy_score(y_test_6, y_pred_6_b))
print("Precision:",metrics.precision_score(y_test_6, y_pred_6_b))
print("Recall:",metrics.recall_score(y_test_6, y_pred_6_b))Output
[stdout]
Accuracy: 0.6553911205073996
Precision: 0.6556655665566556
Recall: 0.9786535303776683In this experiment step the notebook asks tuned_model_6_b â the hyperparameterâtuned classical classifier you trained earlier with GridSearchCV and timed with start=time.time â to produce a hard class prediction for every example in X_test_6, which is the temporally heldâout test matrix of engineered, normalized feature windows for the “6” window configuration. The predict call maps each test window to a discrete label (the Up/Down movement class) aligned with y_test_6 so that the subsequent evaluation code can build a confusion matrix and compute accuracy, precision and recall. This follows the same predictionâevaluate pattern used elsewhere: the LSTM counterpart tuned_model_6_lstm performs an analogous predict call but on X_test_6_lstm (the sequenceâshaped input for the recurrent model), and other classical experiments such as tuned_model_5_b used the same predict call on their own X_test_5; note that earlier in the notebook you also obtained continuous scores via predict_proba for tuned_model_5_b (y_proba_5_b) when a probabilityâbased ranking was required, whereas this predict invocation returns discrete class labels for direct confusionâmatrix style evaluation.
ROC Curce
In [50] Copy
y_proba_6_b=tuned_model_6_b.predict_proba(X_test_6)[:, 1]
fpr, tpr, _=metrics.roc_curve(y_test_6, y_proba_6_b)
auc=metrics.roc_auc_score(y_test_6, y_proba_6_b)
plt.plot(fpr,tpr,label="AUC="+str(auc))
plt.legend(loc=4)
plt.plot([0, 1], [0, 1], linestyle='--') # plot no skill
plt.title('ROC-Curve')
plt.show()Output
The notebook asks tuned_model_6_b to produce class probability estimates for every example in X_test_6 and extracts the probability assigned to the positive (Up) class into y_proba_6_b; those continuous scores and the heldâout true labels y_test_6 are passed to metrics.roc_curve to compute false positive and true positive rates across decision thresholds, and metrics.roc_auc_score to compute the scalar area under the ROC. The computed fpr/tpr pair is drawn with plt.plot and annotated with a legend that includes the AUC, a diagonal noâskill reference line is added, a title is set, and the figure is rendered with plt.show so you can visually assess discriminatory performance on that temporally heldâout partition. This mirrors the earlier evaluation step you saw for tuned_model_5_b and follows the same pattern used for the other folds (for example the tuned_model_1_b and tuned_model_7_b cells), so it provides a consistent probabilistic evaluation across experiments; recall tuned_model_6_b itself was obtained during the model training and hyperparameter search phase whose runtime was recorded with start=time.time.
LSTM
In [51] Copy
start=time.time()
# number of epochs
epochs=1
# number of units
LSTM_units_6_lstm=400
# number of samples
num_samples=1
# time_steps
look_back=1
# numer of features
num_features_6_lstm=X_train_6.shape[1]
# Regularization
dropout_rate=0.
recurrent_dropout=0.5
# print
verbose=0
#hyperparameter
batch_size=[1]
# hyperparameter
hyperparameter_6_lstm={'batch_size':batch_size}
# create Classifier
clf_6_lstm=KerasClassifier(build_fn=create_shallow_LSTM,
epochs=epochs,
LSTM_units=LSTM_units_6_lstm,
num_samples=num_samples,
look_back=look_back,
num_features=num_features_6_lstm,
dropout_rate=dropout_rate,
recurrent_dropout=recurrent_dropout,
verbose=verbose)
# Gridsearch
search_6_lstm=GridSearchCV(estimator=clf_6_lstm,
param_grid=hyperparameter_6_lstm,
n_jobs=-1,
cv=tscv,
scoring=scoring_lstm, # accuracy
refit=True,
return_train_score=False)
# Fit model
tuned_model_6_lstm=search_6_lstm.fit(X_train_6_lstm, y_train_6, shuffle=False, callbacks=[reset])
print("\n")
# View Accuracy
print(scoring_lstm +' of the best model: ', tuned_model_6_lstm.best_score_)
# best_score_ Mean cross-validated score of the best_estimator
print("\n")
# View best hyperparameters
print("Best hyperparameters:")
print('epochs:', tuned_model_6_lstm.best_estimator_.get_params()['epochs'])
print('batch_size:', tuned_model_6_lstm.best_estimator_.get_params()['batch_size'])
print('dropout_rate:', tuned_model_6_lstm.best_estimator_.get_params()['dropout_rate'])
print('recurrent_dropout:', tuned_model_6_lstm.best_estimator_.get_params()['recurrent_dropout'])
end=time.time()
print("\n")
print("Running Time:", end - start)Output
[stdout]
accuracy of the best model: 0.49563492063492065
Best hyperparameters:
epochs: 1
batch_size: 1
dropout_rate: 0.0
recurrent_dropout: 0.5
Running Time: 674.0648419857025The cell begins by capturing a wallâclock timestamp into the variable start so the notebook can report how long the LSTM experiment takes to run; that timestamp is taken immediately before the expensive work of instantiating a KerasClassifier, wrapping it in a GridSearchCV, and calling the fit routine on the timeâwindowed training arrays X_train_6_lstm and y_train_6 (the actual fit call appears later in the cell). Later the cell captures another timestamp into end and prints the difference as Running Time, producing a simple elapsedâtime measurement for this specific model configuration and hyperparameter search. Within the experiment pipeline that turns engineered, normalized windows into LSTM inputs and compares learners, this timing pattern provides a perâexperiment performance metric useful for benchmarking and reproducibility; the notebook repeats the same pattern for other window configurations (the cells for the 1, 2 and 4 window experiments do the same), so start serves as the consistent start marker for each timing block while end and the subtraction yield the measured runtime for that block.
Confusion Matrix
In [52] Copy
# Make predictions
y_pred_6_lstm=tuned_model_6_lstm.predict(X_test_6_lstm)
# create confustion matrix
fig, ax=plt.subplots()
sns.heatmap(pd.DataFrame(metrics.confusion_matrix(y_test_6, y_pred_6_lstm)), annot=True, cmap="Blues" ,fmt='g')
plt.title('Confusion matrix'); plt.ylabel('Actual label'); plt.xlabel('Predicted label')
ax.xaxis.set_ticklabels(['Down', 'Up']); ax.yaxis.set_ticklabels(['Down', 'Up'])
print("Accuracy:",metrics.accuracy_score(y_test_6, y_pred_6_lstm))
print("Precision:",metrics.precision_score(y_test_6, y_pred_6_lstm))
print("Recall:",metrics.recall_score(y_test_6, y_pred_6_lstm))Output
[stdout]
Accuracy: 0.638477801268499
Precision: 0.6471885336273429
Recall: 0.9638752052545156After hyperparameter tuning and model construction the notebook asks tuned_model_6_lstm to produce class predictions for the temporally heldâout, timeâwindowed test set X_test_6_lstm so we can evaluate how the LSTM trained for the “6” window configuration performs against the true labels y_test_6. The predicted labels are compared to y_test_6 by building a confusion matrix via metrics.confusion_matrix, converting that into a pandas DataFrame and visualizing it with seaborn’s heatmap while annotating cells and labeling the axes as Down and Up so the distribution of true versus predicted directions is clear. Finally, the notebook computes scalar summary metricsâaccuracy, precision and recallâusing metrics.accuracy_score, metrics.precision_score and metrics.recall_score and prints them, providing the numeric performance that will be compared to the classical model results such as those obtained earlier with tuned_model_6_b on X_test_6; the main difference being that tuned_model_6_lstm consumes the 3D LSTM-shaped windows (X_test_6_lstm) produced earlier when num_features was wired into the KerasClassifier.
ROC Curve
In [53] Copy
y_proba_6_b=tuned_model_6_b.predict_proba(X_test_6)[:, 1]
fpr, tpr, _=metrics.roc_curve(y_test_6, y_proba_6_b)
auc=metrics.roc_auc_score(y_test_6, y_proba_6_b)
plt.plot(fpr,tpr,label="AUC="+str(auc))
plt.legend(loc=4)
plt.plot([0, 1], [0, 1], linestyle='--') # plot no skill
plt.title('ROC-Curve')
plt.show()Output
Once tuned_model_6_b has been trained and the temporally held-out test matrix X_test_6 has been prepared by the preprocessing and time-windowing pipeline, the notebook asks the fitted classifier to produce class-probability estimates for every example in X_test_6 and captures the probability the model assigns to the positive (Up) class into y_proba_6_b. Those continuous scores, rather than hard labels, are required by the subsequent ROC/AUC calculation because they let metrics.roc_curve and metrics.roc_auc_score compute false positive and true positive rates across thresholds and a threshold-agnostic performance summary; this step therefore converts the tuned classical learnerâs outputs into the format the evaluation code expects. The pattern mirrors the same probability-extraction used for the other window experiments (e.g., the 1- and 7-window runs), with the only differences being the specific tuned model and the corresponding test set names produced earlier in the notebook.
Model 7: Volatility, Return and Trading Volume
Baseline
In [54] Copy
# Model specific Parameter
# Number of iterations
iterations_7_b=[10]
# Grid Search
# Regularization
alpha_g_7_b=[0.0019, 0.002, 0.0021]
l1_ratio_g_7_b=[0, 0.2, 0.4, 0.6, 0.8, 1]
# Create hyperparameter options
hyperparameters_g_7_b={'logistic__alpha':alpha_g_7_b,
'logistic__l1_ratio':l1_ratio_g_7_b,
'logistic__penalty':penalty_b,
'logistic__max_iter':iterations_7_b}
# Create grid search
search_g_7_b=GridSearchCV(estimator=pipeline_b,
param_grid=hyperparameters_g_7_b,
cv=tscv,
verbose=0,
n_jobs=-1,
scoring=scoring_b,
refit=metric_b,
return_train_score=False)
# Setting refit='Accuracy', refits an estimator on the whole dataset with the parameter setting that has the best cross-validated mean Accuracy score.
# For multiple metric evaluation, this needs to be a string denoting the scorer is used to find the best parameters for refitting the estimator at the end
# If return_train_score=True training results of CV will be saved as well
# Fit grid search
tuned_model_7_b=search_g_7_b.fit(X_train_7, y_train_7)
#search_g_7_b.cv_results_
# Random Search
# Create regularization hyperparameter distribution using uniform distribution
#alpha_r_7_b=uniform(loc=0.00006, scale=0.002) #loc=0.00006, scale=0.002
#l1_ratio_r_7_b=uniform(loc=0, scale=1)
# Create hyperparameter options
#hyperparameters_r_7_b={'logistic__alpha':alpha_r_7_b, 'logistic__l1_ratio':l1_ratio_r_7_b, 'logistic__penalty':penalty_b,'logistic__max_iter':iterations_7_b}
# Create randomized search
#search_r_7_b=RandomizedSearchCV(pipeline_b, hyperparameters_r_7_b, n_iter=10, random_state=1, cv=tscv, verbose=0, n_jobs=-1, scoring=scoring_b, refit=metric_b, return_train_score=False)
# Setting refit='Accuracy', refits an estimator on the whole dataset with the parameter setting that has the best cross-validated Accuracy score.
# Fit randomized search
#tuned_model_7_b=search_r_4_b.fit(X_train_7, y_train_7)
# View Cost function
print('Loss function:', tuned_model_7_b.best_estimator_.get_params()['logistic__loss'])
# View Accuracy
print(metric_b +' of the best model: ', tuned_model_7_b.best_score_);print("\n")
# best_score_ Mean cross-validated score of the best_estimator
# View best hyperparameters
print("Best hyperparameters:")
print('Number of iterations:', tuned_model_7_b.best_estimator_.get_params()['logistic__max_iter'])
print('Penalty:', tuned_model_7_b.best_estimator_.get_params()['logistic__penalty'])
print('Alpha:', tuned_model_7_b.best_estimator_.get_params()['logistic__alpha'])
print('l1_ratio:', tuned_model_7_b.best_estimator_.get_params()['logistic__l1_ratio'])
# Find the number of nonzero coefficients (selected features)
print("Total number of features:", len(tuned_model_7_b.best_estimator_.steps[1][1].coef_[0][:]))
print("Number of selected features:", np.count_nonzero(tuned_model_7_b.best_estimator_.steps[1][1].coef_[0][:]))
# Gridsearch table
plt.title('Gridsearch')
pvt_7_b=pd.pivot_table(pd.DataFrame(tuned_model_7_b.cv_results_), values='mean_test_accuracy', index='param_logistic__l1_ratio', columns='param_logistic__alpha')
ax_7_b=sns.heatmap(pvt_7_b, cmap="Blues")
plt.show()Output
[stdout]
Loss function: log
accuracy of the best model: 0.5227513227513227
Best hyperparameters:
Number of iterations: 10
Penalty: l1
Alpha: 0.0019
l1_ratio: 0
Total number of features: 69
Number of selected features: 11iterations_7_b is the small list that supplies the candidate value for the logistic step’s maximum-iteration hyperparameter used during hyperparameter search for Model 7; that list is placed into the hyperparameter dictionary under the logistic__max_iter key so GridSearchCV will include that value when it enumerates parameter combinations for pipeline_b with time-series cross-validation (tscv) and scoring defined by scoring_b/refit via metric_b. In practice the list contains a single integer, ten, which signals the grid search to evaluate a max-iteration budget of ten for the logistic estimator across all CV folds for the Volatility, Return and Trading Volume feature set; the pattern mirrors the other model blocks (for example iterations_5_b and iterations_4_b) where each model supplies its own single-value list so the grid param_grid accepts it as an iterable, but the concrete integer can differ between models to reflect prior choices about how many solver iterations that feature configuration requires for convergence. The value flows into search_g_7_b and therefore into tuned_model_7_b, which is later used to produce predictions and probability scores in the same way tuned_model_6_b was used earlier.
Confusion Matrix
In [55] Copy
# Make predictions
y_pred_7_b=tuned_model_7_b.predict(X_test_7)
# create confustion matrix
fig, ax=plt.subplots()
sns.heatmap(pd.DataFrame(metrics.confusion_matrix(y_test_7, y_pred_7_b)), annot=True, cmap="Blues" ,fmt='g')
plt.title('Confusion matrix'); plt.ylabel('Actual label'); plt.xlabel('Predicted label')
ax.xaxis.set_ticklabels(['Down', 'Up']); ax.yaxis.set_ticklabels(['Down', 'Up'])
print("Accuracy:",metrics.accuracy_score(y_test_7, y_pred_7_b))
print("Precision:",metrics.precision_score(y_test_7, y_pred_7_b))
print("Recall:",metrics.recall_score(y_test_7, y_pred_7_b))Output
[stdout]
Accuracy: 0.6617336152219874
Precision: 0.656554712892741
Recall: 0.9950738916256158The assignment to y_pred_7_b directs the hyperparameterâtuned classical classifier tuned_model_7_b to produce discrete class predictions for every temporally heldâout example in X_test_7, where X_test_7 comes from the preprocessing pipeline that framed the historical price series into sevenâstep time windows and normalized the engineered features; those predicted labels are then compared against the true labels y_test_7 using sklearn.metrics.confusion_matrix to produce a contingency table and seaborn.heatmap to visualize perâclass errors (the heatmap is annotated and ticked as Down/Up for interpretability), and the same predicted/true pair is passed to sklearn.metrics.accuracy_score, precision_score, and recall_score so the notebook prints overall accuracy plus positiveâclass precision and sensitivity; this follows the identical postâtuning evaluation pattern used for other window configurations (for example y_pred_1_b, y_pred_2_b, and y_pred_4_b), thereby making the “7” window results directly comparable to the other experiments in the endâtoâend pipeline.
ROC Curve
In [56] Copy
y_proba_7_b=tuned_model_7_b.predict_proba(X_test_7)[:, 1]
fpr, tpr, _=metrics.roc_curve(y_test_7, y_proba_7_b)
auc=metrics.roc_auc_score(y_test_7, y_proba_7_b)
plt.plot(fpr,tpr,label="AUC="+str(auc))
plt.legend(loc=4)
plt.plot([0, 1], [0, 1], linestyle='--') # plot no skill
plt.title('ROC-Curve')
plt.show()Output
The statement calls predict_proba on tuned_model_7_b with the held-out test feature set X_test_7 to obtain probabilistic outputs for each test example and stores the probability assigned to the positive class in y_proba_7_b; in the context of this end-to-end experiment notebookâwhere we already obtained discrete predictions earlier via y_pred_7_b and where tuned_model_7_b is the hyperparameterâtuned classical pipeline produced by GridSearchCVâthese probability estimates are the thresholdâagnostic scores the notebook uses next to compute the ROC curve and AUC via sklearn.metrics functions and to plot the classifierâs discrimination performance, following the same pattern applied for the other tuned_model_* instances elsewhere in the project.
LSTM
In [57] Copy
start=time.time()
# number of epochs
epochs=1
# number of units
LSTM_units_7_lstm=220
# numer of features
num_features_7_lstm=X_train_7.shape[1]
# Regularization
dropout_rate=0.
recurrent_dropout=0.4
# print
verbose=0
#hyperparameter
batch_size=[1]
# hyperparameter
hyperparameter_7_lstm={'batch_size':batch_size}
# create Classifier
clf_7_lstm=KerasClassifier(build_fn=create_shallow_LSTM,
epochs=epochs,
LSTM_units=LSTM_units_7_lstm,
num_samples=num_samples,
look_back=look_back,
num_features=num_features_7_lstm,
dropout_rate=dropout_rate,
recurrent_dropout=recurrent_dropout,
verbose=verbose)
# Gridsearch
search_7_lstm=GridSearchCV(estimator=clf_7_lstm,
param_grid=hyperparameter_7_lstm,
n_jobs=-1,
cv=tscv,
scoring=scoring_lstm, # accuracy
refit=True,
return_train_score=False)
# Fit model
tuned_model_7_lstm=search_7_lstm.fit(X_train_7_lstm, y_train_7, shuffle=False, callbacks=[reset])
print("\n")
# View Accuracy
print(scoring_lstm +' of the best model: ', tuned_model_7_lstm.best_score_)
# best_score_ Mean cross-validated score of the best_estimator
print("\n")
# View best hyperparameters
print("Best hyperparameters:")
print('epochs:', tuned_model_7_lstm.best_estimator_.get_params()['epochs'])
print('batch_size:', tuned_model_7_lstm.best_estimator_.get_params()['batch_size'])
print('dropout_rate:', tuned_model_7_lstm.best_estimator_.get_params()['dropout_rate'])
print('recurrent_dropout:', tuned_model_7_lstm.best_estimator_.get_params()['recurrent_dropout'])
end=time.time()
print("\n")
print("Running Time:", end - start)Output
[stdout]
accuracy of the best model: 0.5313492063492063
Best hyperparameters:
epochs: 1
batch_size: 1
dropout_rate: 0.0
recurrent_dropout: 0.4
Running Time: 288.78774094581604At the top of the LSTM training cell the notebook captures a wallâclock timestamp so the experiment can report how long the whole model-search and fit block took to run. Within this endâtoâend experiment, runtime reporting is part of the experimentâs metadata â it sits alongside the crossâvalidated score and the best hyperparameter values as an operational metric that lets you compare compute cost between model variants and between LSTM runs and the classical learners. After recording the initial timestamp, the cell proceeds to declare the model configuration and hyperparameter grid, constructs the KerasClassifier wrapper around create_shallow_LSTM (recall the earlier LSTM search used scoring_lstm and passed num_features into create_shallow_LSTM), builds a GridSearchCV with the timeâseries crossâvalidator tscv and scoring_lstm, and calls fit with shuffle disabled and the reset callback. When fit returns, the notebook captures a second timestamp, computes the difference, and prints the elapsed Running Time together with the tuned_model_7_lstm best score and the best_estimator_ parameters obtained via get_params. This patternârecord start, run model selection and fit, record end, print elapsed timeâappears repeatedly for other LSTM variants in the notebook; the only differences across those cells are the model hyperparameters (units, dropout, number of features) while the timing wrapper and reporting logic remain the same.
Confusion Matrix
In [58] Copy
# Make predictions
y_pred_7_lstm=tuned_model_7_lstm.predict(X_test_7_lstm)
# create confustion matrix
fig, ax=plt.subplots()
sns.heatmap(pd.DataFrame(metrics.confusion_matrix(y_test_7, y_pred_7_lstm)), annot=True, cmap="Blues" ,fmt='g')
plt.title('Confusion matrix'); plt.ylabel('Actual label'); plt.xlabel('Predicted label')
ax.xaxis.set_ticklabels(['Down', 'Up']); ax.yaxis.set_ticklabels(['Down', 'Up'])
print("Accuracy:",metrics.accuracy_score(y_test_7, y_pred_7_lstm))
print("Precision:",metrics.precision_score(y_test_7, y_pred_7_lstm))
print("Recall:",metrics.recall_score(y_test_7, y_pred_7_lstm))Output
[stdout]
Accuracy: 0.6553911205073996
Precision: 0.6539717083786725
Recall: 0.986863711001642Like the earlier step that produced y_pred_7_b from the tuned classical pipeline, the notebook asks the tuned LSTM model tuned_model_7_lstm to produce discrete class predictions for the timeâwindowed test set X_test_7_lstm and stores those labels in y_pred_7_lstm; X_test_7_lstm is the temporally framed, normalized sequence input produced by the preprocessing stage so the LSTM can operate on short sliding windows. The predicted labels are then summarized with a confusion matrix computed from y_test_7 and y_pred_7_lstm, rendered as a seaborn heatmap on a matplotlib figure with the axes annotated as the negative class labeled “Down” and the positive class labeled “Up.” Finally, the notebook prints common classification summariesâaccuracy, precision, and recallâusing sklearn.metrics to quantify how well the tuned LSTM separates Up versus Down on the heldâout temporal test set, enabling a direct comparison of the recurrent modelâs performance with the classical learners evaluated earlier.
ROC Curve
In [59] Copy
y_proba_7_b=tuned_model_7_b.predict_proba(X_test_7)[:, 1]
fpr, tpr, _=metrics.roc_curve(y_test_7, y_proba_7_b)
auc=metrics.roc_auc_score(y_test_7, y_proba_7_b)
plt.plot(fpr,tpr,label="AUC="+str(auc))
plt.legend(loc=4)
plt.plot([0, 1], [0, 1], linestyle='--') # plot no skill
plt.title('ROC-Curve')
plt.show()Output
After you already used tuned_model_7_b to produce discrete predictions for the test fold (y_pred_7_b), the notebook calls tuned_model_7_b.predict_proba with X_test_7 to obtain probabilistic forecasts for each test example; from that two-column probability output it takes the second column as the estimated probability of the positive class for each sample. Those positive-class probabilities are then fed to metrics.roc_curve together with the true labels y_test_7 to compute the false positive rate and true positive rate across all decision thresholds, and metrics.roc_auc_score is used to summarize that curve as a single AUC number. Finally, the pipeline draws the ROC curve using the computed fpr and tpr, annotates it with the AUC in the legend, overlays a diagonal noâskill line for reference, sets the title to indicate an ROC plot, and renders the figure. Within the endâtoâend experiment, this produces a thresholdâindependent measure of tuned_model_7_bâs ability to discriminate future up/down moves on X_test_7, following the same ROC/AUC evaluation pattern used for the other tuned_model_* runs.
Download the source code using the button below: